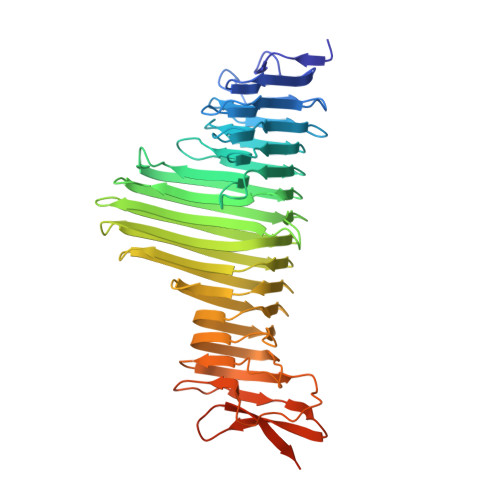
Unique structural features of a bacterial autotransporter adhesin suggest mechanisms for interaction with host macromolecules.
Paxman, J.J., Lo, A.W., Sullivan, M.J., Panjikar, S., Kuiper, M., Whitten, A.E., Wang, G., Luan, C.H., Moriel, D.G., Tan, L., Peters, K.M., Phan, M.D., Gee, C.L., Ulett, G.C., Schembri, M.A., Heras, B.(2019) Nat Commun 10: 1967-1967
- PubMed: 31036849 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-019-09814-6
- Primary Citation Related Structures:
6BEA - PubMed Abstract:
Autotransporters are the largest family of outer membrane and secreted proteins in Gram-negative bacteria. Most autotransporters are localised to the bacterial surface where they promote colonisation of host epithelial surfaces. Here we present the crystal structure of UpaB, an autotransporter that is known to contribute to uropathogenic E. coli (UPEC) colonisation of the urinary tract. We provide evidence that UpaB can interact with glycosaminoglycans and host fibronectin. Unique modifications to its core β-helical structure create a groove on one side of the protein for interaction with glycosaminoglycans, while the opposite face can bind fibronectin. Our findings reveal far greater diversity in the autotransporter β-helix than previously thought, and suggest that this domain can interact with host macromolecules. The relevance of these interactions during infection remains unclear.
- Department of Biochemistry and Genetics, La Trobe Institute for Molecular Science, La Trobe University, Melbourne, 3086, VIC, Australia.
Organizational Affiliation: