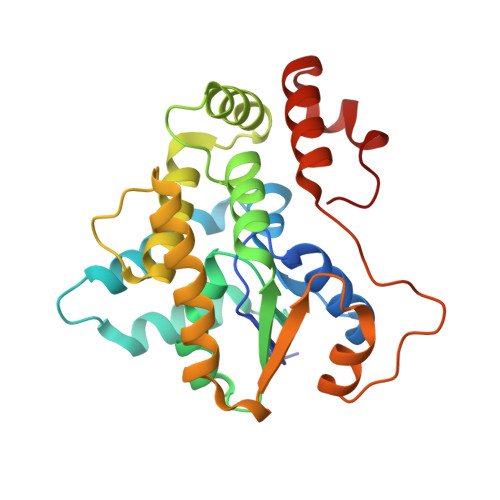
Why does oxamniquine kill Schistosoma mansoni and not S. haematobium and S. japonicum?
Rugel, A.R., Guzman, M.A., Taylor, A.B., Chevalier, F.D., Tarpley, R.S., McHardy, S.F., Cao, X., Holloway, S.P., Anderson, T.J.C., Hart, P.J., LoVerde, P.T.(2020) Int J Parasitol