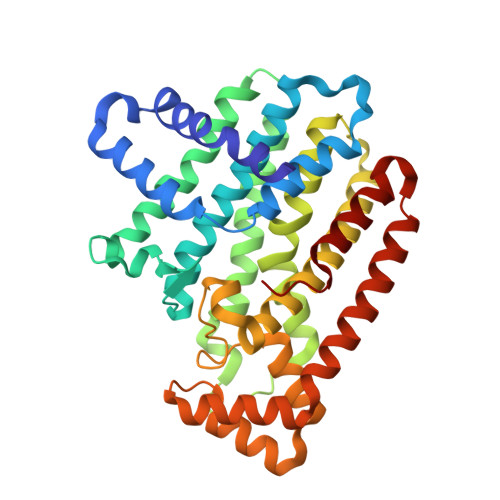
Structural characterization of a lepidopteran type-II farnesyl diphosphate synthase from the spruce budworm, Choristoneura fumiferana: Implications for inhibitor design.
Picard, M.E., Nisole, A., Beliveau, C., Sen, S., Barbar, A., Shi, R., Cusson, M.(2017) Insect Biochem Mol Biol 92: 84-92
- PubMed: 29183817 Search on PubMed
- DOI: https://doi.org/10.1016/j.ibmb.2017.11.011
- Primary Citation Related Structures:
6B02, 6B04, 6B06, 6B07 - PubMed Abstract:
Farnesyl diphosphate synthase (FPPS) is an enzyme from the class of short chain (E)-prenyltransferases that catalyzes the condensation of two molecules of isopentenyl diphosphate (IPP, C 5 ) with dimethylallyl diphosphate (DMAPP, C 5 ) to generate the C 15 product FPP. In insects, FPPS plays a key role in the biosynthesis of the morphogenetic and gonadotropic "juvenile hormone" (JH). Lepidopteran genomes encode two very distinct FPPS paralogs, one of which ("type-II") is expressed almost exclusively in the JH-producing glands, the corpora allata. This paralog has been hypothesized to display structural features that enable the binding of the bulkier precursors required for the biosynthesis of lepidopteran ethyl-branched JHs. Here, we report on the first crystal structures of an insect FPPS solved to date. Apo, ligand-bound, and inhibitor-bound structures of type-II FPPS (FPPS2) from the spruce budworm, Choristoneura fumiferana (Order: Lepidoptera), were obtained. Comparison of apo and inhibitor-bound enzymes revealed differences in both inhibitor binding and structural plasticity of CfFPPS2 compared to other FPPSs. Our data showed that IPP is not essential to the closure of the C-terminal tail. Ortho-substituted pyridinium bisphosphonates, previously shown to inhibit CfFPPS2, bound to the allylic site, as predicted; however, their alkyl groups were oriented towards the homoallylic binding site, with the bulkier propyl-substituted inhibitor penetrating deeply into the IPP binding pocket. The current study sheds light on the structural basis of substrate specificity of type-II FPPS of the spruce budworm. Through a comparison with other inhibitor-bound FPPSs, we propose several approaches to improve inhibitor selectivity and potency.
- Département de biochimie, de microbiologie et de bio-informatique, Institut de Biologie Intégrative et des Systèmes, PROTEO, Université Laval, Quebec City, QC, G1V 0A6, Canada. Electronic address: marie-eve.picard.8@ulaval.ca.
Organizational Affiliation: