Identification of selective, brain penetrant Toxoplasma gondii dihydrofolate reductase inhibitors for the treatment of toxoplasmosis
Thomas, S.B., Hopper, A.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Bifunctional dihydrofolate reductase-thymidylate synthase | 566 | Toxoplasma gondii | Mutation(s): 0 EC: 1.5.1.3 (PDB Primary Data), 2.1.1.45 (PDB Primary Data) | 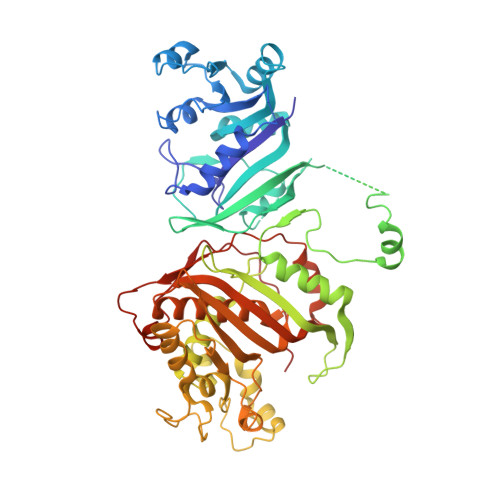 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q07422 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 4 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| NDP Download:Ideal Coordinates CCD File | D [auth A], H [auth B] | NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE C21 H30 N7 O17 P3 ACFIXJIJDZMPPO-NNYOXOHSSA-N |  | ||
| CB3 Download:Ideal Coordinates CCD File | E [auth A], I [auth B] | 10-PROPARGYL-5,8-DIDEAZAFOLIC ACID C24 H23 N5 O6 LTKHPMDRMUCUEB-IBGZPJMESA-N |  | ||
| UMP Download:Ideal Coordinates CCD File | C [auth A], G [auth B] | 2'-DEOXYURIDINE 5'-MONOPHOSPHATE C9 H13 N2 O8 P JSRLJPSBLDHEIO-SHYZEUOFSA-N |  | ||
| CP6 Download:Ideal Coordinates CCD File | F [auth A], J [auth B] | 5-(4-CHLORO-PHENYL)-6-ETHYL-PYRIMIDINE-2,4-DIAMINE C12 H13 Cl N4 WKSAUQYGYAYLPV-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 53.589 | α = 90 |
| b = 144.916 | β = 90 |
| c = 176.044 | γ = 90 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| HKL-2000 | data reduction |
| HKL-2000 | data scaling |
| PHASER | phasing |
| HKL-2000 | data collection |
| HKL-2000 | data extraction |
| HKL-2000 | data processing |