Crystal structure of DMF5 TCR bound to HLA-A2 presenting synthetic peptide MMWDRGLGMM
Riley, T.P., Baker, B.M.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| HLA class I histocompatibility antigen, A-2 alpha chain | 273 | Homo sapiens | Mutation(s): 0 Gene Names: HLA-A, HLAA | 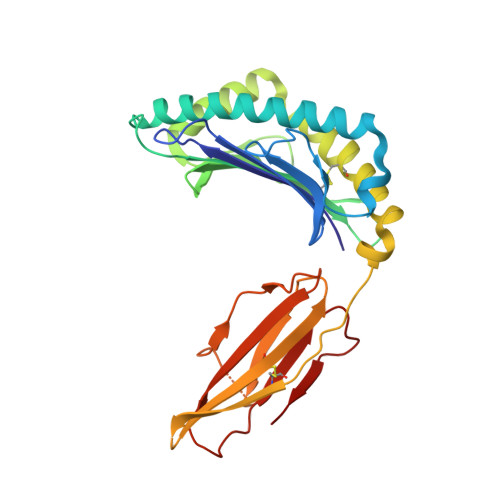 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P04439 GTEx: ENSG00000206503 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P04439 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Beta-2-microglobulin | 100 | Homo sapiens | Mutation(s): 0 Gene Names: B2M, CDABP0092, HDCMA22P | 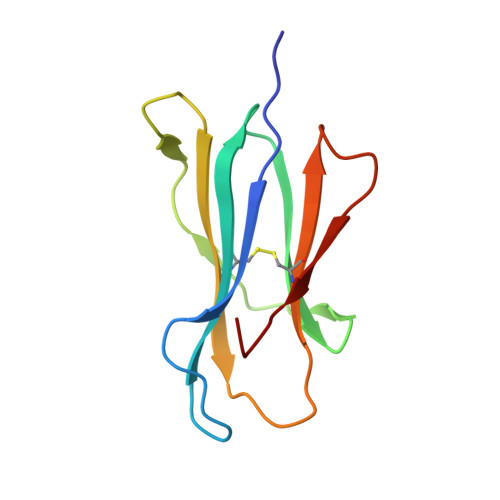 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P61769 GTEx: ENSG00000166710 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P61769 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| MET-MET-TRP-ASP-ARG-GLY-LEU-GLY-MET-MET | 10 | synthetic construct | Mutation(s): 0 |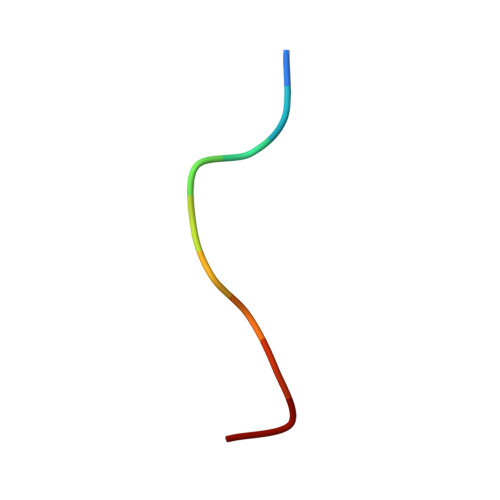  | |
Entity ID: 4 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| DMF5 TCR alpha chain | 197 | Homo sapiens | Mutation(s): 0 | 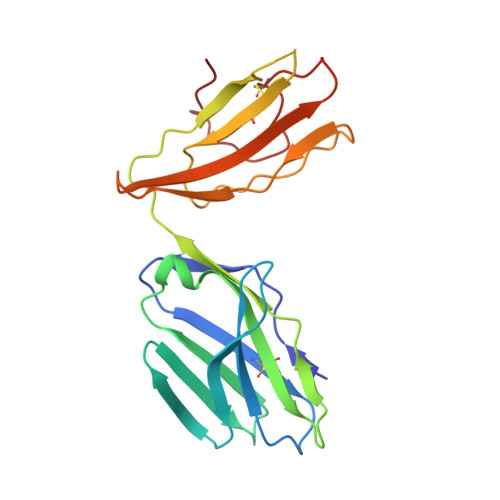 | |
Entity ID: 5 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| DMF5 TCR beta chain | 241 | Homo sapiens | Mutation(s): 0 | 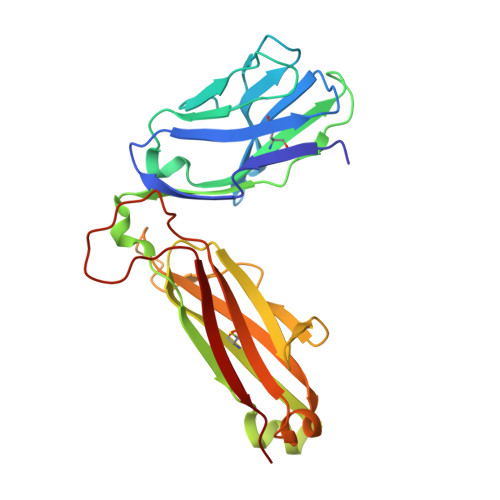 | |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 232.078 | α = 90 |
| b = 50.326 | β = 99.52 |
| c = 84.361 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| HKL-2000 | data scaling |
| PDB_EXTRACT | data extraction |
| HKL-2000 | data reduction |
| PHASER | phasing |