Optimization of Chromeno[2,3- c]pyrrol-9(2 H)-ones as Highly Potent, Selective, and Orally Bioavailable PDE5 Inhibitors: Structure-Activity Relationship, X-ray Crystal Structure, and Pharmacodynamic Effect on Pulmonary Arterial Hypertension.
Wu, D., Huang, Y., Chen, Y., Huang, Y.Y., Geng, H., Zhang, T., Zhang, C., Li, Z., Guo, L., Chen, J., Luo, H.B.(2018) J Med Chem 61: 8468-8473
- PubMed: 30148362 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.8b01209
- Primary Citation Related Structures:
5ZZ2, 6ACB - PubMed Abstract:
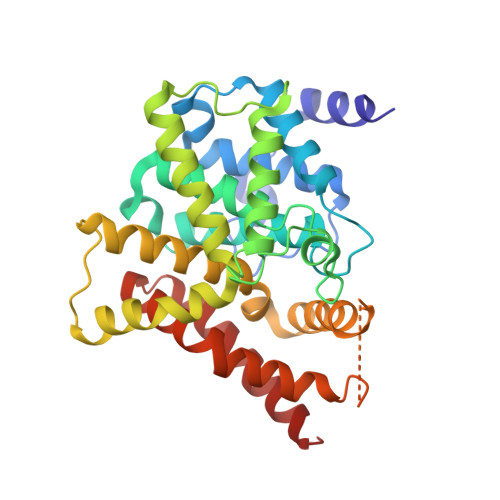
To further explore the structure-activity relationship around the chromeno[2,3- c]pyrrol-9(2 H)-one scaffold, 19 derivatives as inhibitors against PDE5 were discovered. The most potent inhibitor 3 has an IC 50 of 0.32 nM with remarkable selectivity and druglike profile. Oral administration of 3 (1.25 mg/kg) caused comparable therapeutic effects to sildenafil (10.0 mg/kg) against pulmonary arterial hypertension. Further, different binding patterns from sildenafil were revealed in cocrystal structures, which provide structural templates for discovery of highly potent PDE5 inhibitors.
- School of Pharmaceutical Sciences , Sun Yat-Sen University , Guangzhou 510006 , P. R. China.
Organizational Affiliation: