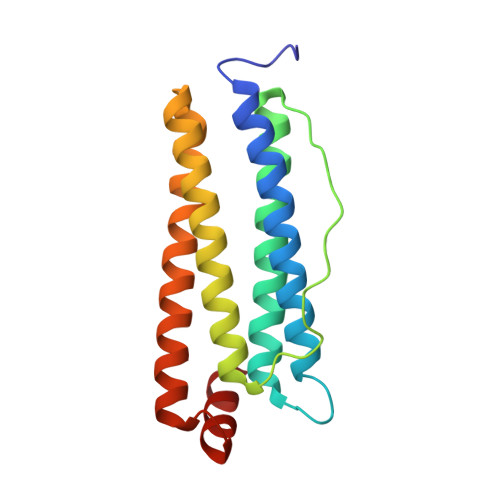
The first crystal structure of crustacean ferritin that is a hybrid type of H and L ferritin
Masuda, T., Zang, J., Zhao, G., Mikami, B.(2018) Protein Sci 27: 1955-1960
- PubMed: 30099791 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.3495
- Primary Citation Related Structures:
6A4U - PubMed Abstract:
Ferritin, a ubiquitous iron storage protein, has a crucial role in innate immunity in arthropods, which have no adaptive immune system. Arthropods are thought to have two types of ferritin molecules: the secreted type and the cytosolic type. Here, we present the first crystal structure of ferritin from crustacean, kuruma prawn (Marsupenaeus japonicus), at 1.16 Å resolution. This shrimp ferritin (MjFer) is the cytosolic type, and its structure shows well-conserved ferritin fold composed of a 4-helix bundle that assembles into a cage-like 24-mer. The structure of MjFer was more similar to those of human and vertebrate ferritins than to that of the secreted-type arthropod ferritin from an insect. MjFer possesses both a ferroxidase site and a nucleation site, which are the main characteristics of vertebrate H and L chain ferritins, respectively. The first crystal structure of crustacean ferritin, MjFer, has exceptionally high quality that provides the detailed structural information of metal moving pathway in ferritin.
- Laboratory of Food Quality Design and Development, Division of Agronomy and Horticultural Science, Graduate School of Agriculture, Kyoto University, Kyoto, 611-0011, Japan.
Organizational Affiliation: