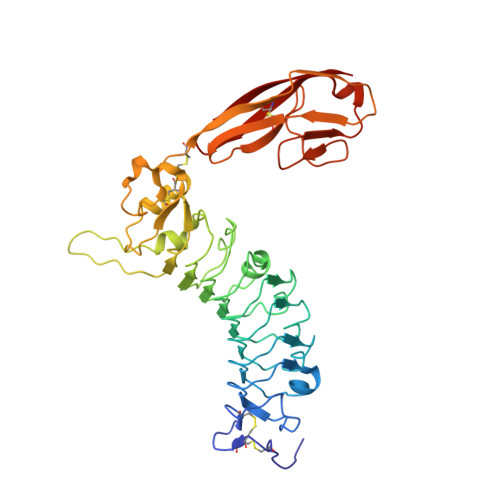
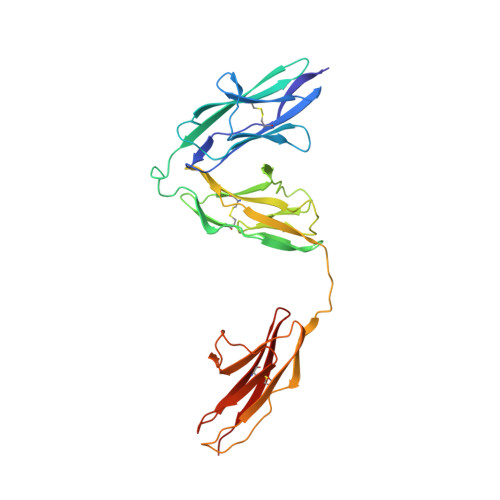Structural basis of SALM5-induced PTP delta dimerization for synaptic differentiation
Lin, Z., Liu, J., Ding, H., Xu, F., Liu, H.(2018) Nat Commun 9: 268-268
- PubMed: 29348579 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-017-02414-2
- Primary Citation Related Structures:
5XNP, 5XNQ - PubMed Abstract:
SALM5, a synaptic adhesion molecule implicated in autism, induces presynaptic differentiation through binding to the LAR family receptor protein tyrosine phosphatases (LAR-RPTPs) that have been highlighted as presynaptic hubs for synapse formation. The mechanisms underlying SALM5/LAR-RPTP interaction remain unsolved. Here we report crystal structures of human SALM5 LRR-Ig alone and in complex with human PTPδ Ig1-3 (MeA - ). Distinct from other LAR-RPTP ligands, SALM5 mainly exists as a dimer with LRR domains from two protomers packed in an antiparallel fashion. In the 2:2 heterotetrameric SALM5/PTPδ complex, a SALM5 dimer bridges two separate PTPδ molecules. Structure-guided mutations and heterologous synapse formation assays demonstrate that dimerization of SALM5 is prerequisite for its functionality in inducing synaptic differentiation. This study presents a structural template for the SALM family and reveals a mechanism for how a synaptic adhesion molecule directly induces cis-dimerization of LAR-RPTPs into higher-order signaling assembly.
- State Key Laboratory of Natural and Biomimetic Drugs & Department of Molecular and Cellular Pharmacology, School of Pharmaceutical Sciences, Peking University Health Science Center, 38 Xueyuan Road, Haidian District, Beijing, 100191, China.
Organizational Affiliation: