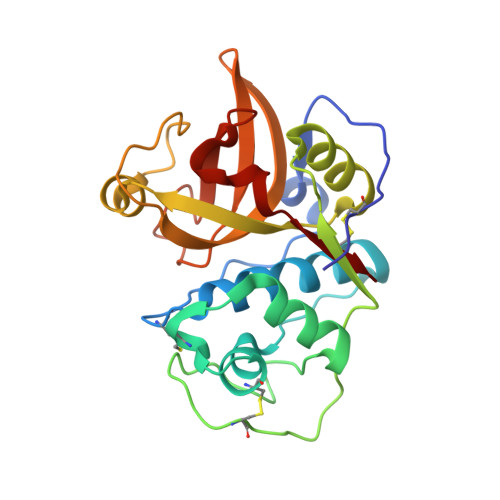
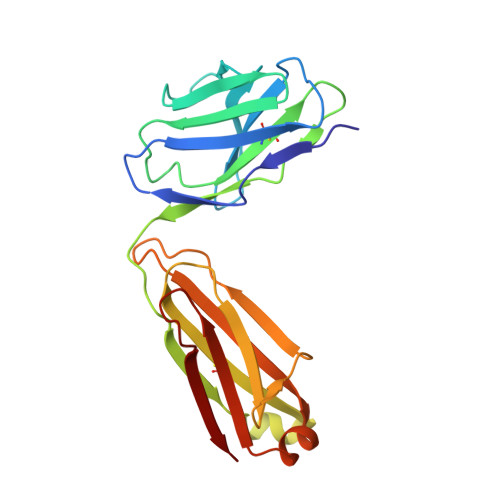
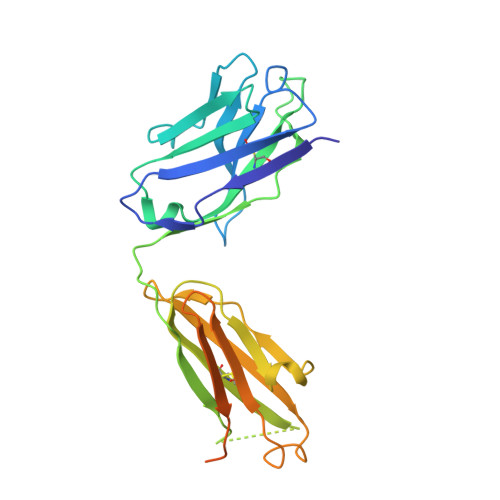
Structural Analysis of Der p 1-Antibody Complexes and Comparison with Complexes of Proteins or Peptides with Monoclonal Antibodies.
Osinski, T., Pomes, A., Majorek, K.A., Glesner, J., Offermann, L.R., Vailes, L.D., Chapman, M.D., Minor, W., Chruszcz, M.(2015) J Immunol 195: 307-316
- PubMed: 26026055 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.4049/jimmunol.1402199
- Primary Citation Related Structures:
4POZ, 5VCN, 5VCO - PubMed Abstract:
Der p 1 is a major allergen from the house dust mite, Dermatophagoides pteronyssinus, that belongs to the papain-like cysteine protease family. To investigate the antigenic determinants of Der p 1, we determined two crystal structures of Der p 1 in complex with the Fab fragments of mAbs 5H8 or 10B9. Epitopes for these two Der p 1-specific Abs are located in different, nonoverlapping parts of the Der p 1 molecule. Nevertheless, surface area and identity of the amino acid residues involved in hydrogen bonds between allergen and Ab are similar. The epitope for mAb 10B9 only showed a partial overlap with the previously reported epitope for mAb 4C1, a cross-reactive mAb that binds Der p 1 and its homolog Der f 1 from Dermatophagoides farinae. Upon binding to Der p 1, the Fab fragment of mAb 10B9 was found to form a very rare α helix in its third CDR of the H chain. To provide an overview of the surface properties of the interfaces formed by the complexes of Der p 1-10B9 and Der p 1-5H8, along with the complexes of 4C1 with Der p 1 and Der f 1, a broad analysis of the surfaces and hydrogen bonds of all complexes of Fab-protein or Fab-peptide was performed. This work provides detailed insight into the cross-reactive and specific allergen-Ab interactions in group 1 mite allergens. The surface data of Fab-protein and Fab-peptide interfaces can be used in the design of conformational epitopes with reduced Ab binding for immunotherapy.
- University of Virginia, Charlottesville, VA 22908; Adam Mickiewicz University, 61-712 Poznan, Poland;
Organizational Affiliation: