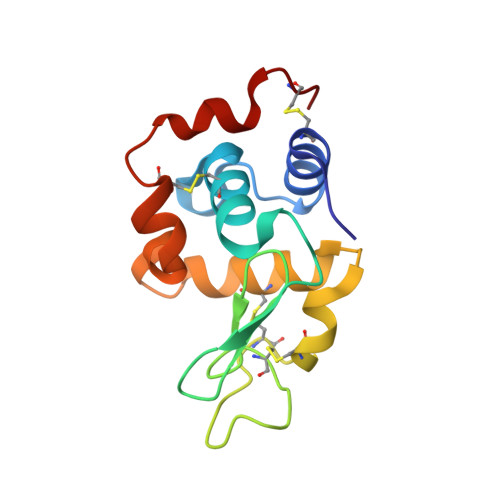
The metalation of hen egg white lysozyme impacts protein stability as shown by ion mobility mass spectrometry, differential scanning calorimetry, and X-ray crystallography.
Sullivan, M.P., Groessl, M., Meier, S.M., Kingston, R.L., Goldstone, D.C., Hartinger, C.G.(2017) Chem Commun (Camb) 53: 4246-4249
- PubMed: 28361137 Search on PubMed
- DOI: https://doi.org/10.1039/c6cc10150j
- Primary Citation Related Structures:
5V4G, 5V4H, 5V4I - PubMed Abstract:
Metalation of hen egg white lysozyme (HEWL) with organometallics was studied with physicochemical methods in solid state, solution and the gas phase. While metalation did not affect the crystal structure of HEWL significantly, protein destabilisation was detected in gas phase and solution.
- School of Chemical Sciences, University of Auckland, Private Bag 92019, Auckland, 1142, New Zealand. c.hartinger@auckland.ac.nz.
Organizational Affiliation: