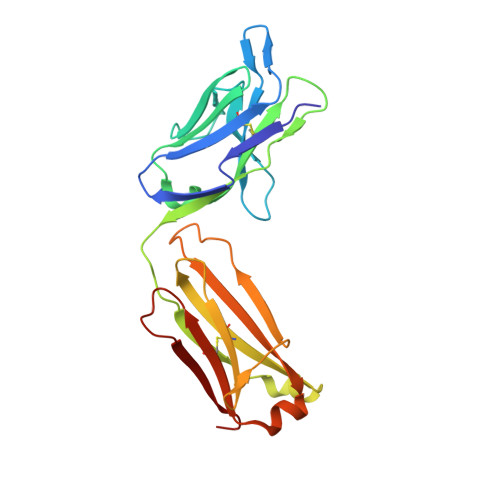
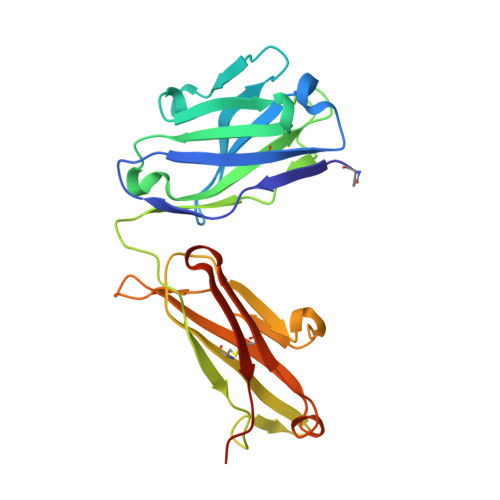
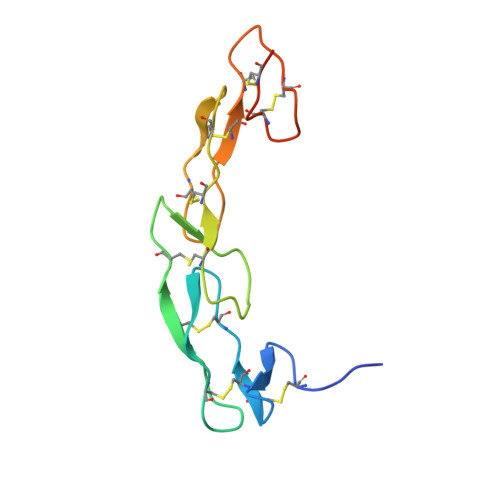
Crystal structure of CD27 in complex with a neutralizing noncompeting antibody.
Teplyakov, A., Obmolova, G., Malia, T.J., Gilliland, G.L.(2017) Acta Crystallogr F Struct Biol Commun 73: 294-299
- PubMed: 28471362 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X17005957
- Primary Citation Related Structures:
5TL5 - PubMed Abstract:
CD27 is a T-cell and B-cell co-stimulatory glycoprotein of the tumor necrosis factor (TNF) receptor superfamily that is dependent on the availability of the TNF-like ligand CD70. Therapeutic approaches to treating autoimmune diseases and cancers with antagonistic and agonistic anti-CD27 monoclonal antibodies (mAbs), respectively, have recently been developed. Mouse anti-human CD27 mAb 2177 shows potency in neutralizing CD70-induced signaling; however, it does not block the binding of soluble CD70. To provide insight into the mechanism of action of the mAb, the crystal structure of the CD27 extracellular domain in complex with the Fab fragment of mAb 2177 was determined at 1.8 Å resolution. CD27 exhibits the assembly of cysteine-rich domains characteristic of the TNF receptor superfamily. The structure reveals a unique binding site of mAb 2177 at the edge of the receptor molecule, which allows the mAb to sterically block the cell-bound form of CD70 from reaching CD27 while leaving the ligand epitope clear. This mode of action suggests a potential dual use of mAb 2177 either as an antagonist or as an agonist.
- Janssen Research and Development LLC, 1400 McKean Road, Spring House, PA 19477, USA.
Organizational Affiliation: