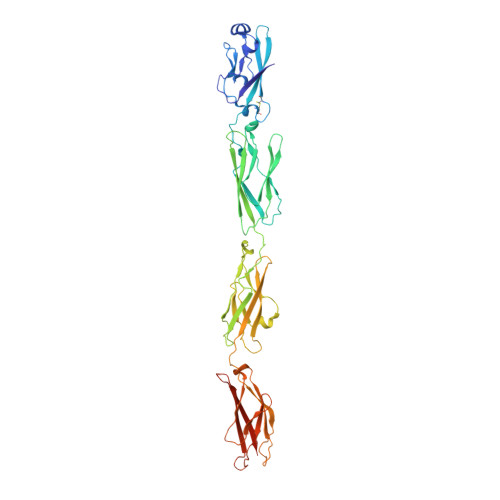
gamma-Protocadherin structural diversity and functional implications.
Goodman, K.M., Rubinstein, R., Thu, C.A., Mannepalli, S., Bahna, F., Ahlsen, G., Rittenhouse, C., Maniatis, T., Honig, B., Shapiro, L.(2016) Elife 5
- PubMed: 27782885 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.20930
- Primary Citation Related Structures:
5SZL, 5SZM, 5SZN, 5SZO, 5SZP, 5SZQ, 5SZR, 5T9T - PubMed Abstract:
Stochastic cell-surface expression of α-, β-, and γ-clustered protocadherins (Pcdhs) provides vertebrate neurons with single-cell identities that underlie neuronal self-recognition. Here we report crystal structures of ectodomain fragments comprising cell-cell recognition regions of mouse γ-Pcdhs γA1, γA8, γB2, and γB7 revealing trans -homodimers, and of C-terminal ectodomain fragments from γ-Pcdhs γA4 and γB2, which depict cis -interacting regions in monomeric form. Together these structures span the entire γ-Pcdh ectodomain. The trans -dimer structures reveal determinants of γ-Pcdh isoform-specific homophilic recognition. We identified and structurally mapped cis -dimerization mutations to the C-terminal ectodomain structures. Biophysical studies showed that Pcdh ectodomains from γB-subfamily isoforms formed cis dimers, whereas γA isoforms did not, but both γA and γB isoforms could interact in cis with α-Pcdhs. Together, these data show how interaction specificity is distributed over all domains of the γ-Pcdh trans interface, and suggest that subfamily- or isoform-specific cis -interactions may play a role in the Pcdh-mediated neuronal self-recognition code.
- Department of Biochemistry and Molecular Biophysics, Columbia University, New York, United States.
Organizational Affiliation: