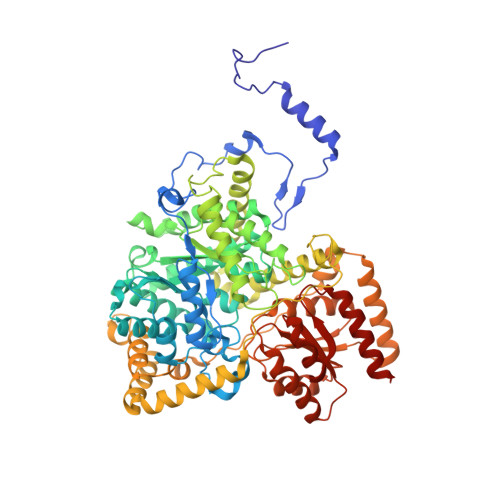
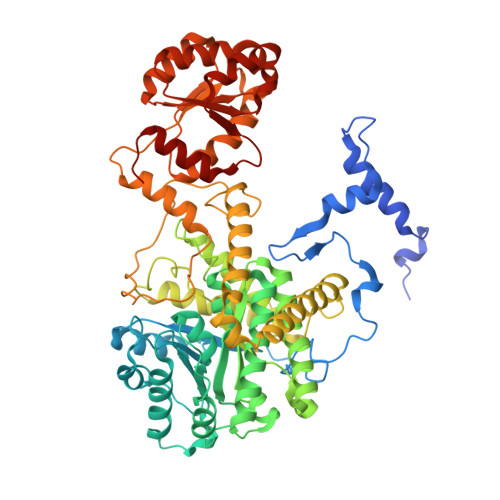
Stabilization of radical intermediates by an active-site tyrosine residue in methylmalonyl-CoA mutase.
Thoma, N.H., Meier, T.W., Evans, P.R., Leadlay, P.F.(1998) Biochemistry 37: 14386-14393
- PubMed: 9772164 Search on PubMed
- DOI: https://doi.org/10.1021/bi981375o
- Primary Citation Related Structures:
5REQ - PubMed Abstract:
The adenosylcobalamin-dependent methylmalonyl-CoA mutase catalyzes the reversible rearrangement of methylmalonyl-CoA into succinyl-CoA by a free-radical mechanism. The recently solved X-ray crystal structure of methylmalonyl-CoA mutase from Propionibacterium shermanii has shown that tyrosine 89 is an active-site residue involved in substrate binding. The role of tyrosine 89, a conserved residue among methylmalonyl-CoA mutases, has been investigated by using site-directed mutagenesis to replace this residue with phenylalanine. The crystal structure of the Tyr89Phe mutant was determined to 2.2 A resolution and was found to be essentially superimposable on that of wild-type. Mutant and wild-type enzyme have very similar KM values, but kcat for the Tyr89Phe mutant is 580-fold lower than for wild-type. The rate of release of tritium from 5'-[3H]adenosylcobalamin during the enzymatic reaction and its rate of appearance in substrate and product were measured. The tritium released was found to partition unequally between methylmalonyl-CoA and succinyl-CoA, in a ratio of 40:60 when the reaction was initiated by addition of methylmalonyl-CoA and in a ratio of 10:90 when the reaction was initiated by addition of succinyl-CoA. The overall release of tritium was four times faster when succinyl-CoA was used as substrate. The tritium isotope effect on the enzyme catalyzed hydrogen transfer, measured with methylmalonyl-CoA as a substrate, was kH/kT = 30, which is within the expected range for a full primary kinetic tritium isotope effect. The different partitioning of tritium, dependent upon which substrate was used, and the normal value for the kinetic tritium isotope effect contrast markedly with the behavior of wild-type mutase. It appears that the loss of a single interaction involving the hydroxyl group of tyrosine 89 both affects the stability of radical intermediates and decreases the rate of interconversion of the substrate- and product-derived radicals.
- Department of Biochemistry, Cambridge Centre for Molecular Recognition, University of Cambridge, United Kingdom.
Organizational Affiliation: