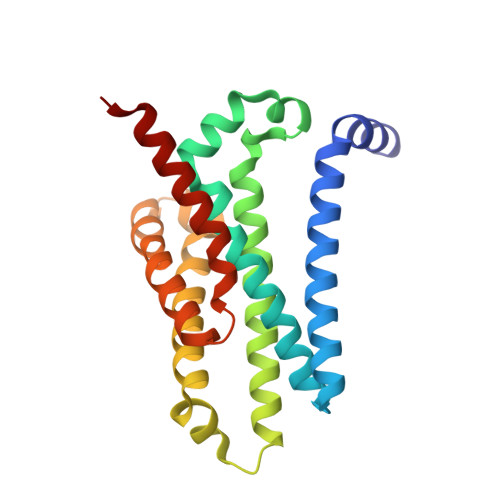
Crystal structures and atomic model of NADPH oxidase.
Magnani, F., Nenci, S., Millana Fananas, E., Ceccon, M., Romero, E., Fraaije, M.W., Mattevi, A.(2017) Proc Natl Acad Sci U S A 114: 6764-6769
- PubMed: 28607049 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1702293114
- Primary Citation Related Structures:
5O0T, 5O0X - PubMed Abstract:
NADPH oxidases (NOXs) are the only enzymes exclusively dedicated to reactive oxygen species (ROS) generation. Dysregulation of these polytopic membrane proteins impacts the redox signaling cascades that control cell proliferation and death. We describe the atomic crystal structures of the catalytic flavin adenine dinucleotide (FAD)- and heme-binding domains of Cylindrospermum stagnale NOX5. The two domains form the core subunit that is common to all seven members of the NOX family. The domain structures were then docked in silico to provide a generic model for the NOX family. A linear arrangement of cofactors (NADPH, FAD, and two membrane-embedded heme moieties) injects electrons from the intracellular side across the membrane to a specific oxygen-binding cavity on the extracytoplasmic side. The overall spatial organization of critical interactions is revealed between the intracellular loops on the transmembrane domain and the NADPH-oxidizing dehydrogenase domain. In particular, the C terminus functions as a toggle switch, which affects access of the NADPH substrate to the enzyme. The essence of this mechanistic model is that the regulatory cues conformationally gate NADPH-binding, implicitly providing a handle for activating/deactivating the very first step in the redox chain. Such insight provides a framework to the discovery of much needed drugs that selectively target the distinct members of the NOX family and interfere with ROS signaling.
- Department of Biology and Biotechnology "L. Spallanzani," University of Pavia, 27100 Pavia, Italy; francesca.magnani@unipv.it andrea.mattevi@unipv.it.
Organizational Affiliation: