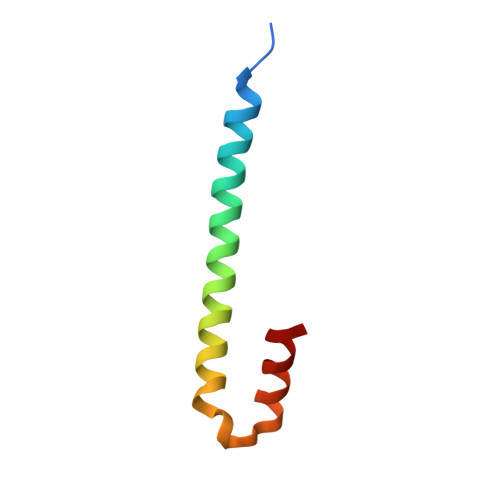
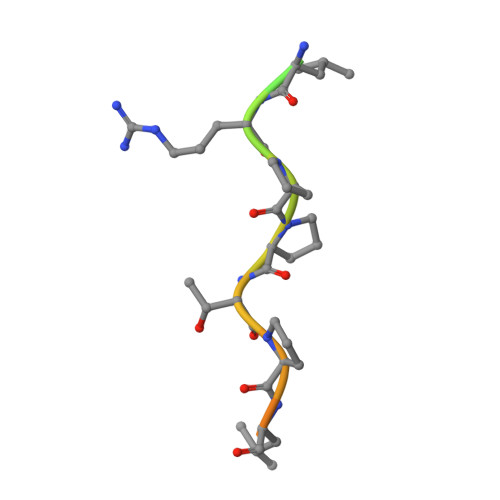
Short Linear Sequence Motif LxxPTPh Targets Diverse Proteins to Growing Microtubule Ends.
Kumar, A., Manatschal, C., Rai, A., Grigoriev, I., Degen, M.S., Jaussi, R., Kretzschmar, I., Prota, A.E., Volkmer, R., Kammerer, R.A., Akhmanova, A., Steinmetz, M.O.(2017) Structure 25: 924-932.e4
- PubMed: 28552577 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2017.04.010
- Primary Citation Related Structures:
5N74 - PubMed Abstract:
Microtubule plus-end tracking proteins (+TIPs) are involved in virtually all microtubule-based processes. End-binding (EB) proteins are considered master regulators of +TIP interaction networks, since they autonomously track growing microtubule ends and recruit a plethora of proteins to this location. Two major EB-interacting elements have been described: CAP-Gly domains and linear SxIP sequence motifs. Here, we identified LxxPTPh as a third EB-binding motif that enables major +TIPs to interact with EBs at microtubule ends. In contrast to EB-SxIP and EB-CAP-Gly, the EB-LxxPTPh binding mode does not depend on the C-terminal tail region of EB. Our study reveals that +TIPs developed additional strategies besides CAP-Gly and SxIP to target EBs at growing microtubule ends. They further provide a unique basis to discover novel +TIPs, and to dissect the role of key interaction nodes and their differential regulation for hierarchical +TIP network organization and function in eukaryotic organisms.
- Laboratory of Biomolecular Research, Division of Biology and Chemistry, Paul Scherrer Institut, 5232 Villigen PSI, Switzerland.
Organizational Affiliation: