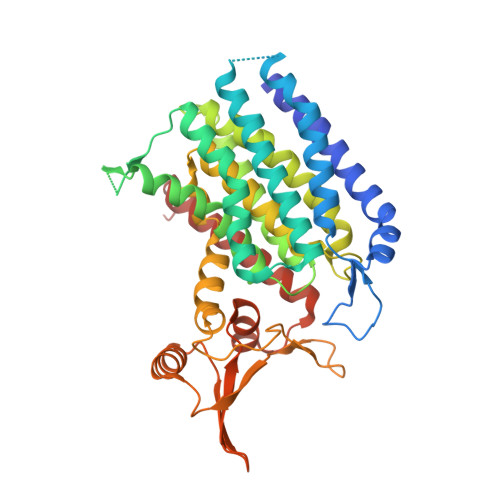
Structures of DPAGT1 Explain Glycosylation Disease Mechanisms and Advance TB Antibiotic Design.
Dong, Y.Y., Wang, H., Pike, A.C.W., Cochrane, S.A., Hamedzadeh, S., Wyszynski, F.J., Bushell, S.R., Royer, S.F., Widdick, D.A., Sajid, A., Boshoff, H.I., Park, Y., Lucas, R., Liu, W.M., Lee, S.S., Machida, T., Minall, L., Mehmood, S., Belaya, K., Liu, W.W., Chu, A., Shrestha, L., Mukhopadhyay, S.M.M., Strain-Damerell, C., Chalk, R., Burgess-Brown, N.A., Bibb, M.J., Barry Iii, C.E., Robinson, C.V., Beeson, D., Davis, B.G., Carpenter, E.P.(2018) Cell 175: 1045-1058.e16
- PubMed: 30388443 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2018.10.037
- Primary Citation Related Structures:
5LEV, 5O5E, 6FM9, 6FWZ - PubMed Abstract:
Protein N-glycosylation is a widespread post-translational modification. The first committed step in this process is catalysed by dolichyl-phosphate N-acetylglucosamine-phosphotransferase DPAGT1 (GPT/E.C. 2.7.8.15). Missense DPAGT1 variants cause congenital myasthenic syndrome and disorders of glycosylation. In addition, naturally-occurring bactericidal nucleoside analogues such as tunicamycin are toxic to eukaryotes due to DPAGT1 inhibition, preventing their clinical use. Our structures of DPAGT1 with the substrate UDP-GlcNAc and tunicamycin reveal substrate binding modes, suggest a mechanism of catalysis, provide an understanding of how mutations modulate activity (thus causing disease) and allow design of non-toxic "lipid-altered" tunicamycins. The structure-tuned activity of these analogues against several bacterial targets allowed the design of potent antibiotics for Mycobacterium tuberculosis, enabling treatment in vitro, in cellulo and in vivo, providing a promising new class of antimicrobial drug.
- Structural Genomics Consortium, University of Oxford, Oxford, OX3 7DQ, UK.
Organizational Affiliation: