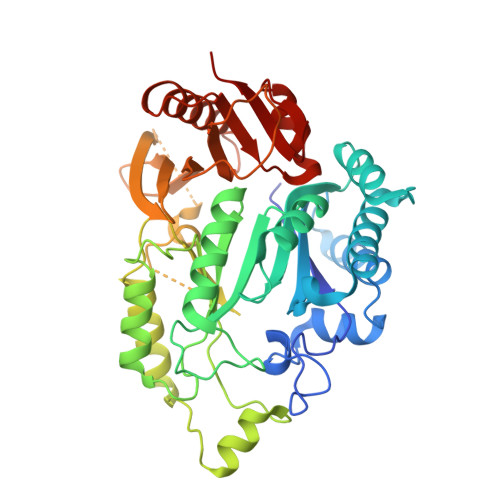
Structural basis for tRNA modification by Elp3 from Dehalococcoides mccartyi.
Glatt, S., Zabel, R., Kolaj-Robin, O., Onuma, O.F., Baudin, F., Graziadei, A., Taverniti, V., Lin, T.Y., Baymann, F., Seraphin, B., Breunig, K.D., Muller, C.W.(2016) Nat Struct Mol Biol 23: 794-802
- PubMed: 27455459 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.3265
- Primary Citation Related Structures:
5L7J, 5L7L - PubMed Abstract:
During translation elongation, decoding is based on the recognition of codons by corresponding tRNA anticodon triplets. Molecular mechanisms that regulate global protein synthesis via specific base modifications in tRNA anticodons are receiving increasing attention. The conserved eukaryotic Elongator complex specifically modifies uridines located in the wobble base position of tRNAs. Mutations in Elongator subunits are associated with certain neurodegenerative diseases and cancer. Here we present the crystal structure of D. mccartyi Elp3 (DmcElp3) at 2.15-Å resolution. Our results reveal an unexpected arrangement of Elp3 lysine acetyltransferase (KAT) and radical S-adenosyl methionine (SAM) domains, which share a large interface and form a composite active site and tRNA-binding pocket, with an iron-sulfur cluster located in the dimerization interface of two DmcElp3 molecules. Structure-guided mutagenesis studies of yeast Elp3 confirmed the relevance of our findings for eukaryotic Elp3s and should aid in understanding the cellular functions and pathophysiological roles of Elongator.
- European Molecular Biology Laboratory, Structural and Computational Biology Unit, Heidelberg, Germany.
Organizational Affiliation: