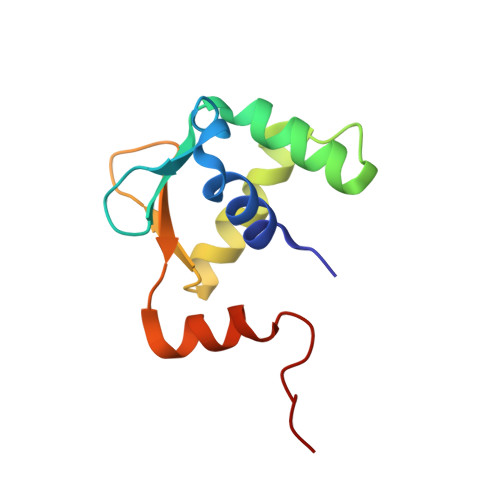
Structured and disordered regions cooperatively mediate DNA-binding autoinhibition of ETS factors ETV1, ETV4 and ETV5.
Currie, S.L., Lau, D.K.W., Doane, J.J., Whitby, F.G., Okon, M., McIntosh, L.P., Graves, B.J.(2017) Nucleic Acids Res 45: 2223-2241
- PubMed: 28161714 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkx068
- Primary Citation Related Structures:
5ILS, 5ILU, 5ILV - PubMed Abstract:
Autoinhibition enables spatial and temporal regulation of cellular processes by coupling protein activity to surrounding conditions, often via protein partnerships or signaling pathways. We report the molecular basis of DNA-binding autoinhibition of ETS transcription factors ETV1, ETV4 and ETV5, which are often overexpressed in prostate cancer. Inhibitory elements that cooperate to repress DNA binding were identified in regions N- and C-terminal of the ETS domain. Crystal structures of these three factors revealed an α-helix in the C-terminal inhibitory domain that packs against the ETS domain and perturbs the conformation of its DNA-recognition helix. Nuclear magnetic resonance spectroscopy demonstrated that the N-terminal inhibitory domain (NID) is intrinsically disordered, yet utilizes transient intramolecular interactions with the DNA-recognition helix of the ETS domain to mediate autoinhibition. Acetylation of selected lysines within the NID activates DNA binding. This investigation revealed a distinctive mechanism for DNA-binding autoinhibition in the ETV1/4/5 subfamily involving a network of intramolecular interactions not present in other ETS factors. These distinguishing inhibitory elements provide a platform through which cellular triggers, such as protein-protein interactions or post-translational modifications, may specifically regulate the function of these oncogenic proteins.
- Department of Oncological Sciences, University of Utah, Salt Lake City, UT 84112-5550, USA.
Organizational Affiliation: