Crystal Structure of chromodomain of CBX7 in complex with inhibitor UNC3866
Liu, Y., Tempel, W., Walker, J.R., Stuckey, J.I., Dickson, B.M., James, L.I., Frye, S.V., Bountra, C., Arrowsmith, C.H., Edwards, A.M., Min, J.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Chromobox protein homolog 7 | 56 | Homo sapiens | Mutation(s): 0 Gene Names: CBX7 | 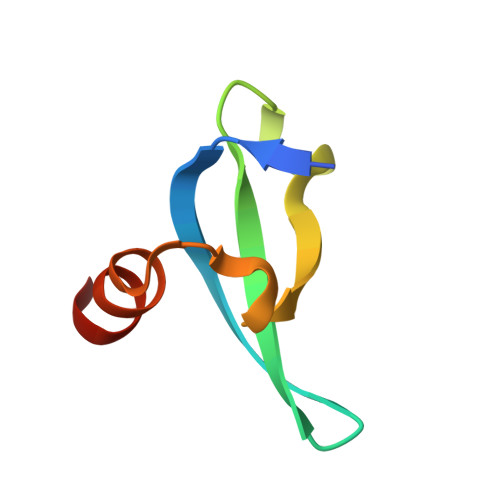 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: O95931 GTEx: ENSG00000100307 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | O95931 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| peptide-like inhibitor UNC3866 | 6 | Homo sapiens | Mutation(s): 0 | 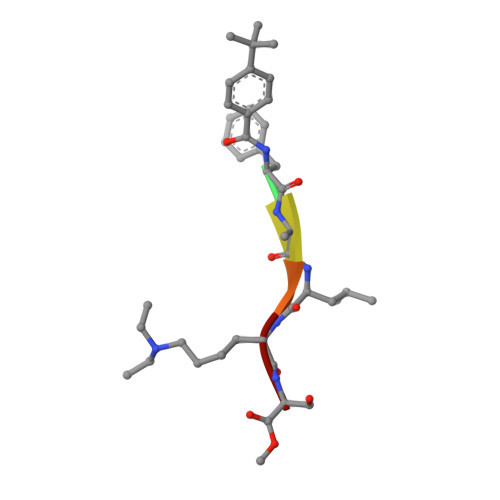 | |
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| UNX Download:Ideal Coordinates CCD File | C [auth A] D [auth A] E [auth A] F [auth A] G [auth A] | UNKNOWN ATOM OR ION X |  | ||
| Modified Residues 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| 5R5 Query on 5R5 | B | L-PEPTIDE LINKING | C4 H9 N O3 |  | SER |
| ELY Query on ELY | B | L-PEPTIDE LINKING | C10 H22 N2 O2 |  | LYS |
| Entity ID: 2 | |||||
|---|---|---|---|---|---|
| ID | Chains | Name | Type/Class | 2D Diagram | 3D Interactions |
| PRD_002208 Query on PRD_002208 | B | UNC3866 | Oligopeptide / Inhibitor |  | |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 40.286 | α = 90 |
| b = 40.286 | β = 90 |
| c = 82.092 | γ = 90 |
| Software Name | Purpose |
|---|---|
| Aimless | data scaling |
| REFMAC | refinement |
| PDB_EXTRACT | data extraction |
| PHASER | phasing |