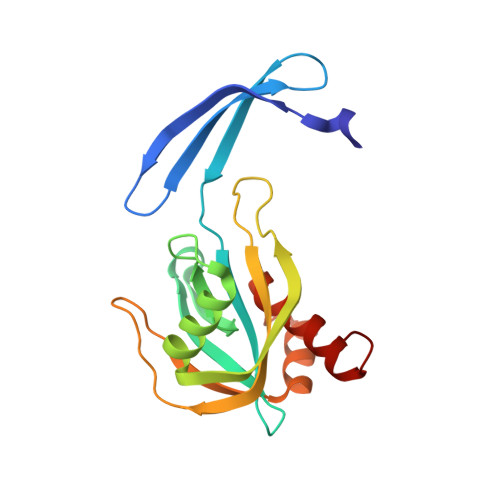
Structural and Enzymatic Characterization of a Nucleoside Diphosphate Sugar Hydrolase from Bdellovibrio bacteriovorus.
de la Pena, A.H., Suarez, A., Duong-Ly, K.C., Schoeffield, A.J., Pizarro-Dupuy, M.A., Zarr, M., Pineiro, S.A., Amzel, L.M., Gabelli, S.B.(2015) PLoS One 10: e0141716-e0141716
- PubMed: 26524597 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0141716
- Primary Citation Related Structures:
5C7Q, 5C7T, 5C8L - PubMed Abstract:
Given the broad range of substrates hydrolyzed by Nudix (nucleoside diphosphate linked to X) enzymes, identification of sequence and structural elements that correctly predict a Nudix substrate or characterize a family is key to correctly annotate the myriad of Nudix enzymes. Here, we present the structure determination and characterization of Bd3179 -- a Nudix hydrolase from Bdellovibrio bacteriovorus-that we show localized in the periplasmic space of this obligate Gram-negative predator. We demonstrate that the enzyme is a nucleoside diphosphate sugar hydrolase (NDPSase) and has a high degree of sequence and structural similarity to a canonical ADP-ribose hydrolase and to a nucleoside diphosphate sugar hydrolase (1.4 and 1.3 Å Cα RMSD respectively). Examination of the structural elements conserved in both types of enzymes confirms that an aspartate-X-lysine motif on the C-terminal helix of the α-β-α NDPSase fold differentiates NDPSases from ADPRases.
- Department of Biomedical Engineering, Johns Hopkins University School of Medicine, Baltimore, Maryland, United States of America; Structural Enzymology and Thermodynamics Group, Department of Biophysics and Biophysical Chemistry, Johns Hopkins University School of Medicine, Baltimore, Maryland, United States of America.
Organizational Affiliation: