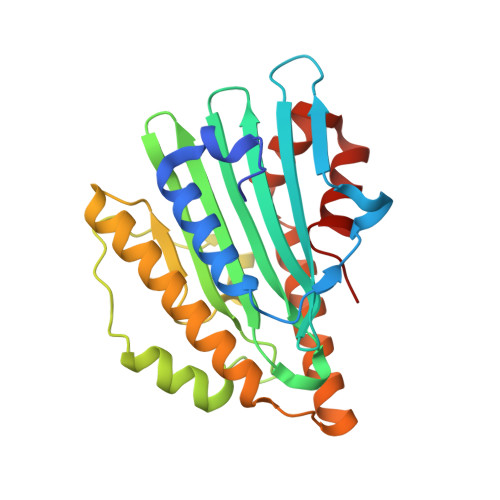
Atomic-resolution structure of the phycocyanobilin:ferredoxin oxidoreductase I86D mutant in complex with fully protonated biliverdin
Hagiwara, Y., Wada, K., Irikawa, T., Sato, H., Unno, M., Yamamoto, K., Fukuyama, K., Sugishima, M.(2016) FEBS Lett 590: 3425-3434
- PubMed: 27596987 Search on PubMed
- DOI: https://doi.org/10.1002/1873-3468.12387
- Primary Citation Related Structures:
5B4H, 5B4I, 5B4J - PubMed Abstract:
Phycocyanobilin:ferredoxin oxidoreductase (PcyA) catalyzes the reduction of biliverdin (BV) to produce phycocyanobilin, a linear tetrapyrrole pigment used for light harvesting and light sensing. Spectroscopic and HPLC analyses inidicate that BV bound to the I86D mutant of PcyA is fully protonated (BVH + ) and can accept an electron, but I86D is unable to donate protons for the reduction; therefore, compared to the wild-type PcyA, the I86D mutant stabilizes BVH + . To elucidate the structural basis of the I86D mutation, we determined the atomic-resolution structure of the I86D-BVH + complex and the protonation states of the essential residues Asp105 and Glu76 in PcyA. Our study revealed that Asp105 adopted a fixed conformation in the I86D mutant, although it had dual conformations in wild-type PcyA which reflected the protonation states of BV. Taken together with biochemical/spectroscopic results, our analysis of the I86D-BVH + structure supports the hypothesis that flexibility of Asp105 is essential for the catalytic activity of PcyA.
- Department of Biochemistry and Applied Chemistry, National Institute of Technology, Kurume College, Kurume, Japan.
Organizational Affiliation: