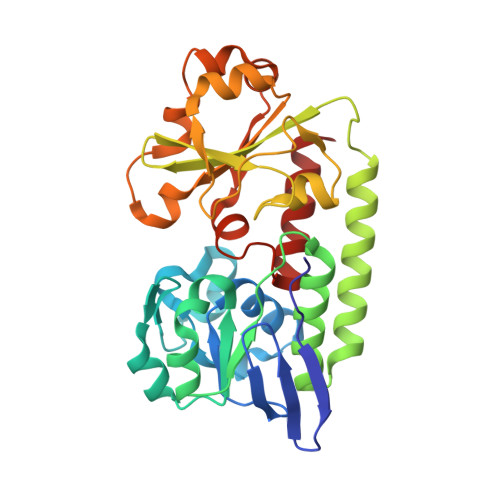
Interactions of the periplasmic binding protein CeuE with Fe(III) n-LICAM(4-) siderophore analogues of varied linker length.
Wilde, E.J., Hughes, A., Blagova, E.V., Moroz, O.V., Thomas, R.P., Turkenburg, J.P., Raines, D.J., Duhme-Klair, A.K., Wilson, K.S.(2017) Sci Rep 7: 45941-45941
- PubMed: 28383577 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep45941
- Primary Citation Related Structures:
5A5D, 5A5V, 5AD1, 5LWH, 5LWQ, 5MBQ, 5MBT, 5MBU, 5TCY - PubMed Abstract:
Bacteria use siderophores to mediate the transport of essential Fe(III) into the cell. In Campylobacter jejuni the periplasmic binding protein CeuE, an integral part of the Fe(III) transport system, has adapted to bind tetradentate siderophores using a His and a Tyr side chain to complete the Fe(III) coordination. A series of tetradentate siderophore mimics was synthesized in which the length of the linker between the two iron-binding catecholamide units was increased from four carbon atoms (4-LICAM 4- ) to five, six and eight (5-, 6-, 8-LICAM 4- , respectively). Co-crystal structures with CeuE showed that the inter-planar angles between the iron-binding catecholamide units in the 5-, 6- and 8-LICAM 4- structures are very similar (111°, 110° and 110°) and allow for an optimum fit into the binding pocket of CeuE, the inter-planar angle in the structure of 4-LICAM 4- is significantly smaller (97°) due to restrictions imposed by the shorter linker. Accordingly, the protein-binding affinity was found to be slightly higher for 5- compared to 4-LICAM 4- but decreases for 6- and 8-LICAM 4- . The optimum linker length of five matches that present in natural siderophores such as enterobactin and azotochelin. Site-directed mutagenesis was used to investigate the relative importance of the Fe(III)-coordinating residues H227 and Y288.
- Structural Biology Laboratory, Department of Chemistry, University of York, Heslington, York YO10 5DD, UK.
Organizational Affiliation: