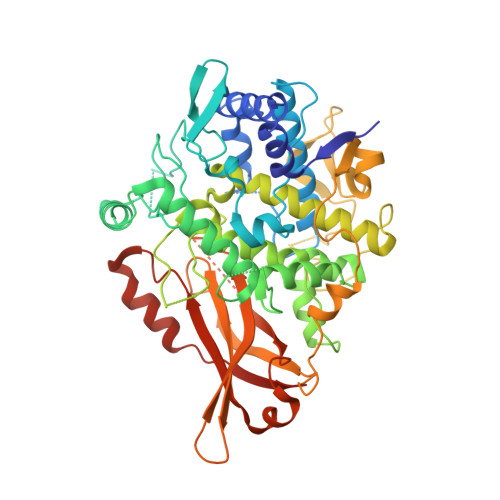
Structural Basis for Highly Efficient Production of Catechol Derivatives at Acidic pH by Tyrosinase from Burkholderia thailandensis
Son, H.F., Lee, S.H., Lee, S.H., Kim, H., Hong, H., Lee, U.J., Lee, P.G., Kim, B.G., Kim, K.J.(2018) ACS Catal 8: 10375-10382