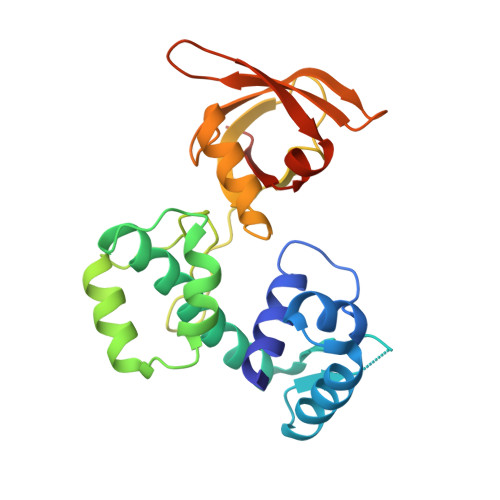
Crystal structures of manganese-dependent transcriptional repressor MntR (Rv2788) from Mycobacterium tuberculosis in apo and manganese bound forms.
Cong, X.Y., Yuan, Z.L., Wang, Z., Wei, B., Xu, S.J., Wang, J.B.(2018) Biochem Biophys Res Commun 501: 423-427
- PubMed: 29730293 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2018.05.005
- Primary Citation Related Structures:
5ZR4, 5ZR6 - PubMed Abstract:
The pathogenic Mycobacterium tuberculosis encodes two members of the DtxR family metalloregulators, IdeR and MntR. IdeR represses gene expression in response to ferrous iron, while MntR (Rv2788) functions as a manganese-dependent transcriptional repressor, which represses the expression of manganese transporter genes to maintain manganese homeostasis. Although the structural study towards IdeR is in-depth, there is no MntR structure available. Herein, we report both apo and manganese bound forms of MntR structures from M. tuberculosis. MntR has evolved into two metal ion binding sites like other DtxR proteins and for the first time, we captured the two sites fully occupied by its natural ions with one Mn 2+ ion at the first site and two Mn 2+ ions at the second binding site (binuclear manganese cluster). The conformation change of MntR resulting from manganese binding could prime the MntR for DNA binding, which is a conserved activation mechanism among DtxR family.
- State Key Laboratory of Microbial Technology, Shandong University, Jinan 250100, China.
Organizational Affiliation: