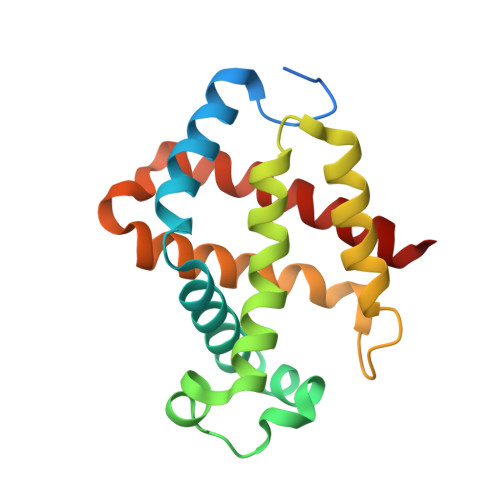
Crystal structure of Kumaglobin: a hexacoordinated heme protein from an anhydrobiotic tardigrade, Ramazzottius varieornatus.
Kim, J., Fukuda, Y., Inoue, T.(2019) FEBS J 286: 1287-1304
- PubMed: 30506636 Search on PubMed
- DOI: https://doi.org/10.1111/febs.14713
- Primary Citation Related Structures:
5ZIQ, 5ZM9 - PubMed Abstract:
Tardigrades, also known as water bears, can survive extreme conditions. For example, tardigrades have high tolerance to extreme desiccation because they can enter an anhydrobiotic state, in which they show no or nearly undetectable metabolic processes. Proteins from anhydrobiotic tardigrades with low homology to known proteins from other organisms are new potential targets for structural genomics. Here, we present spectroscopic and structural characterization of an unprecedented globin protein (Kumaglobin: Kgb) found in an anhydrobiotic tardigrade. Spectroscopy reveals that Kgb contains hexacoordinated low-spin heme, which is not capable of binding to hydrogen sulfide (H 2 S) unlike other globin proteins, such as neuroglobin. Interestingly however, when distal histidine is replaced with alanine, H 2 S is capable of binding to heme, implying that the distal histidine of Kgb binds tightly to heme. The overall structure of Kgb at 1.5 Å resolution shows high resemblance to well-characterized eukaryotic globin proteins, such as myoglobin and cytoglobin. However, the heme coordination geometry in Kgb is unique because the distal histidinyl ligand is located at the 11th position of helix E while it is found at 7th position on helix E in many known globin proteins. The unusual conformation of distal histidine in Kgb is stabilized by a hydrogen bond with the carbonyl O atom of A103. Furthermore, bulky residues exist around the heme cofactor, resulting in a ruffling conformation of the porphyrin ring. Based on our study, Kgb is thought to be involved in electron transfer or enzymatic reactions rather than transporting or storing ligands. DATABASE: Structural data are available in the Protein Data Bank under the accession numbers 5ZIQ (Kgb4-SR) and 5ZM9 (Kgb7-house).
- Department of Applied Chemistry, Graduate School of Engineering, Osaka University, Suita, Japan.
Organizational Affiliation: