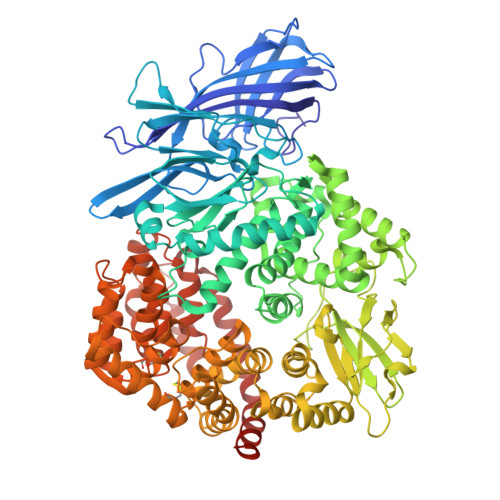
Characterization of the interaction between recombinant porcine aminopeptidase N and spike glycoprotein of porcine epidemic diarrhea virus.
Sun, Y.G., Li, R., Jiang, L., Qiao, S., Zhi, Y., Chen, X.X., Xie, S., Wu, J., Li, X., Deng, R., Zhang, G.(2018) Int J Biol Macromol 117: 704-712
- PubMed: 29802920 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.ijbiomac.2018.05.167
- Primary Citation Related Structures:
5Z65 - PubMed Abstract:
Porcine epidemic diarrhea (PED) has caused huge economic losses to the global pork industry. Infection by its causative agent PED virus (PEDV), an Alpha-coronavirus, was previously proven to be mediated by its spike (S) glycoprotein and a cellular receptor porcine aminopeptidase N (pAPN). Interestingly, some recent studies have indicated that pAPN is not a functional receptor for PEDV. To date, there is a lack of a direct evidence for the interaction between pAPN and PEDV S protein in vitro. Here, we prepared pAPN ectodomain and the truncated variants of PEDV S protein in Drosophila S2 cells. These recombinant proteins were homogeneous after purification by metal-affinity and size-exclusion chromatography. We then assayed the purified target proteins through immunogenicity tests, PEDV binding interference assays, circular dichroism (CD) measurements, pAPN activity assay and structural determination, demonstrating that they were biologically functional. Finally, we characterized their interactions by gel filtration chromatography, native-polyacrylamide gel electrophoresis (PAGE) and surface plasmon resonance (SPR) analyses. The results showed that their affinities were too low to form complexes, which suggest that pAPN may be controversial as the genuine receptor for PEDV. Therefore, further research needs to be carried out to elucidate the interaction between PEDV and its genuine receptor.
- College of Veterinary Medicine, Jilin University, Changchun 130062, Jilin, China; Key Laboratory of Animal Immunology of the Ministry of Agriculture, Henan Provincial Key Laboratory of Animal Immunology, Henan Academy of Agricultural Sciences, Zhengzhou 450002, Henan, China.
Organizational Affiliation: