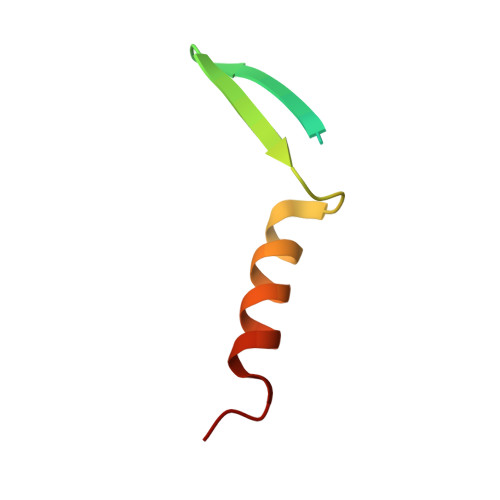
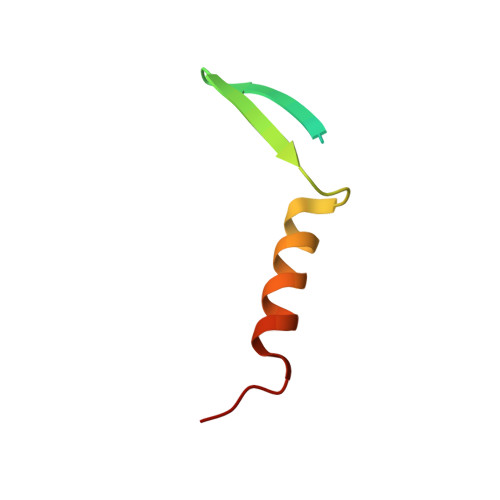
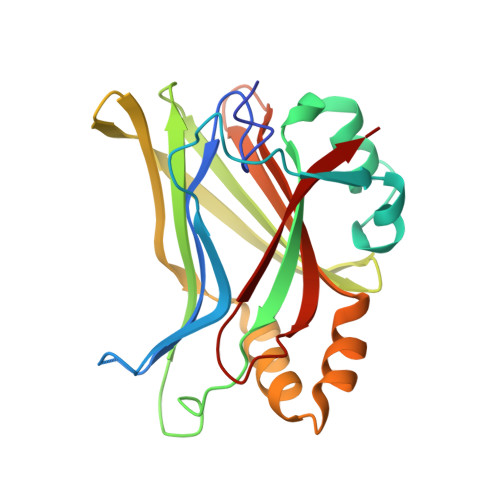
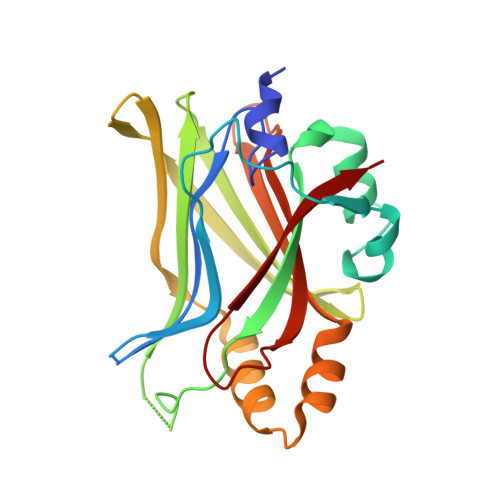
Structural and functional similarity between the Vgll1-TEAD and the YAP-TEAD complexes.
Pobbati, A.V., Chan, S.W., Lee, I., Song, H., Hong, W.(2012) Structure 20: 1135-1140
- PubMed: 22632831 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2012.04.004
- Primary Citation Related Structures:
5Z2Q - PubMed Abstract:
The structure of the complex between the transcription cofactor Vgll1 and the transcription factor TEAD4, the mammalian equivalent of the Drosophila Vestigial and Scalloped, respectively, is determined in this study. Remarkably, Vgll1 interacts with TEAD in a manner similar to the transcription coactivators, as well as oncogenes YAP and TAZ, despite having a varied primary sequence. Vgll1-TEAD complex upregulates the expression of IGFBP-5, a proliferation-promoting gene, and facilitates anchorage-independent cell proliferation. The YAP/TAZ-TEAD complex also upregulates several other proliferation-promoting genes and also promotes anchorage-independent cell proliferation. Given its structural and functional similarity to YAP/TAZ, Vgll1 has the potential to promote cancer progression.
- Cell Biology in Health and Disease Division, Institute of Molecular and Cell Biology, Proteos, Singapore 138673. ajaybabuvp@imcb.a-star.edu.sg
Organizational Affiliation: