Crystal structure of a novel KAS III from Acinetobacter baumannii
Lee, W.C., Jung, M., Lee, J., Kim, Y.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| 3-Oxoacyl-[acyl-carrier-(ACP)] synthase III C terminal family protein | 368 | Acinetobacter baumannii | Mutation(s): 0 EC: 2.3.1.180 (PDB Primary Data), 2.3.1.41 (PDB Primary Data) | 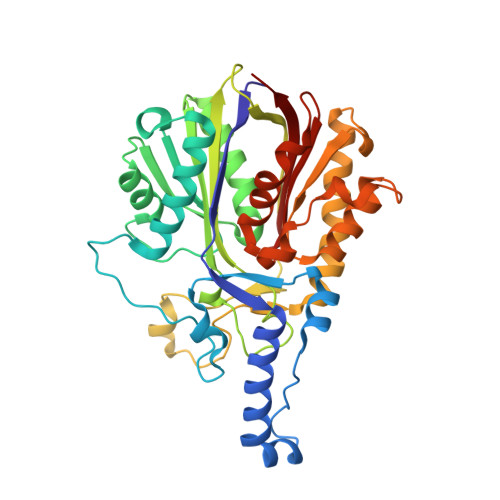 | |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 120.004 | α = 90 |
| b = 88.395 | β = 116.48 |
| c = 77.613 | γ = 90 |
| Software Name | Purpose |
|---|---|
| SCALEPACK | data scaling |
| PHENIX | refinement |
| PDB_EXTRACT | data extraction |
| HKL-2000 | data reduction |
| MOLREP | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Korea, Republic Of | -- |