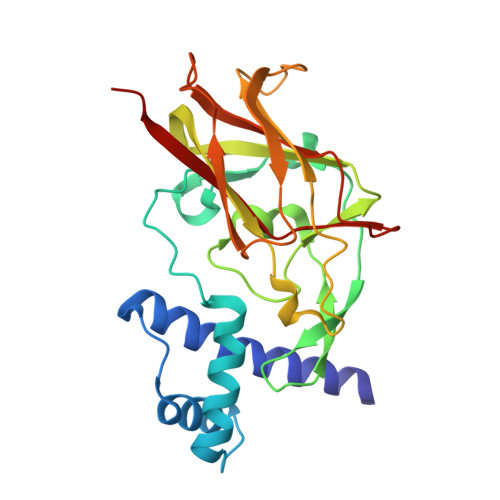
Structural conservation of the autoinhibitory domain in SUN proteins
Xu, Y., Li, W., Ke, H., Feng, W.(2018) Biochem Biophys Res Commun 496: 1337-1343
- PubMed: 29408528 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2018.02.015
- Primary Citation Related Structures:
5YWZ - PubMed Abstract:
LINC complexes span across the nuclear envelope and are assembled by SUN and KASH proteins. SUN1 and SUN2 are the two most abundant SUN proteins in mammals. In SUN2, the predicted coiled-coil domain preceding the SUN domain forms a three-helix bundle that constitutes an autoinhibitory domain (AID) to lock down the SUN domain. Here, we found that SUN1 also contains an AID preceding the SUN domain and solved the structure of the AID-SUN tandem of SUN1. SUN1 AID also adopts a three-helix bundle conformation that interacts with the SUN domain and keeps it in an autoinhibited state. Disruptions of the interaction interface in the AID-SUN tandem restored the SUN domain activity for binding to the KASH peptide. Structural comparison further demonstrated that the autoinhibited conformations of the AID-SUN tandems from SUN1 and SUN2 are similar and the intramolecular interdomain packing in SUN1 is slightly more compact than that in SUN2 due to minor variations of the residues in the interaction interface. Thus, AID is a conserved functional domain in SUN proteins and this work provides the structural evidence to support the conversation of the AID-mediated autoinhibition of SUN proteins.
- National Laboratory of Biomacromolecules, CAS Center for Excellence in Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, 15 Datun Road, Beijing 100101, China; University of Chinese Academy of Sciences, Beijing 100049, China.
Organizational Affiliation: