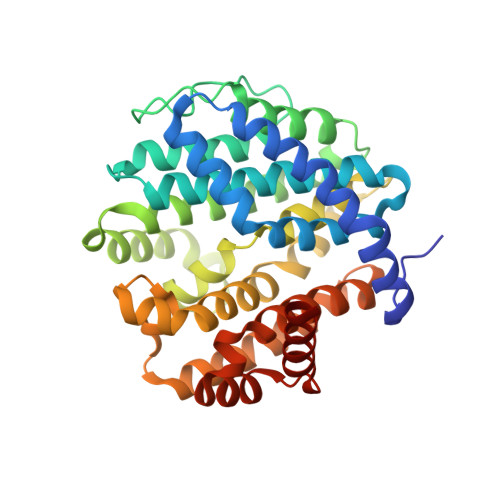
Crystal structure and functional analysis of large-terpene synthases belonging to a newly found subclass.
Fujihashi, M., Sato, T., Tanaka, Y., Yamamoto, D., Nishi, T., Ueda, D., Murakami, M., Yasuno, Y., Sekihara, A., Fuku, K., Shinada, T., Miki, K.(2018) Chem Sci 9: 3754-3758
- PubMed: 29780507 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1039/c8sc00289d
- Primary Citation Related Structures:
5YO8 - PubMed Abstract:
Thousands of terpenes have been identified to date. However, only two classes of enzymes are known to be involved in their biosynthesis, and each class has characteristic amino-acid motifs. We recently identified a novel large-terpene (C 25 /C 30 /C 35 ) synthase, which shares no motifs with known enzymes. To elucidate the molecular mechanism of this enzyme, we determined the crystal structure of a large-β-prene synthase from B. alcalophilus (BalTS). Surprisingly, the overall structure of BalTS is similar to that of the α-domain of class I terpene synthases although their primary structures are totally different from each other. Two novel aspartate-rich motifs, DYLDNLxD and DY(F,L,W)IDxxED, are identified, and mutations of any one of the aspartates eliminate its enzymatic activity. The present work leads us to propose a new subclass of terpene synthases, class IB, which is probably responsible for large-terpene biosynthesis.
- Department of Chemistry , Graduate School of Science , Kyoto University , Sakyo-ku , Kyoto 606-8502 , Japan . Email: mfuji@kuchem.kyoto-u.ac.jp ; Email: miki@kuchem.kyoto-u.ac.jp.
Organizational Affiliation: