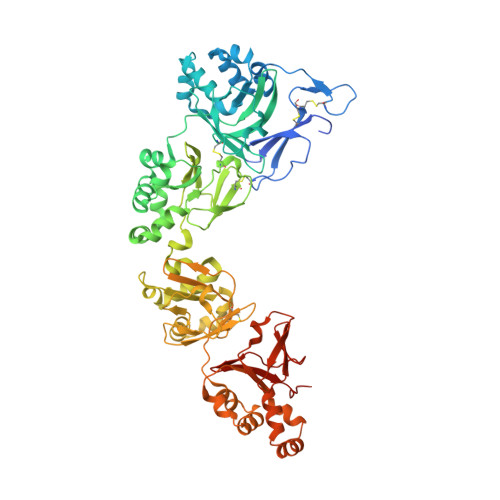
Structural characterizations of human periostin dimerization and cysteinylation.
Liu, J., Zhang, J., Xu, F., Lin, Z., Li, Z., Liu, H.(2018) FEBS Lett 592: 1789-1803
- PubMed: 29754429 Search on PubMed
- DOI: https://doi.org/10.1002/1873-3468.13091
- Primary Citation Related Structures:
5YJG, 5YJH - PubMed Abstract:
Human periostin plays a multifaceted role in remodeling the extracellular matrix milieu by interacting with other proteins and itself in both a heterophilic and homophilic manner. However, the structural mechanism for its extensive interactions has remained elusive. Here, we report the crystal structures of human periostin (EMI-Fas1 I- IV ) and its Cys60Ala mutant. In combination with multi-angle light-scattering analysis and biochemical assays, the crystal structures reveal that periostin mainly exists as a dimer in solution and its homophilic interaction is mainly mediated by the EMI domain. Furthermore, Cys60 undergoes cysteinylation as confirmed by mass spectroscopy, and this site hardly affects the homophilic interaction. Also, the structures yield insights into how periostin forms heterophilic interactions with other proteins under physiological or pathological conditions.
- State Key Laboratory of Natural and Biomimetic Drugs, School of Pharmaceutical Sciences, Peking University Health Science Center, Haidian District, Beijing, China.
Organizational Affiliation: