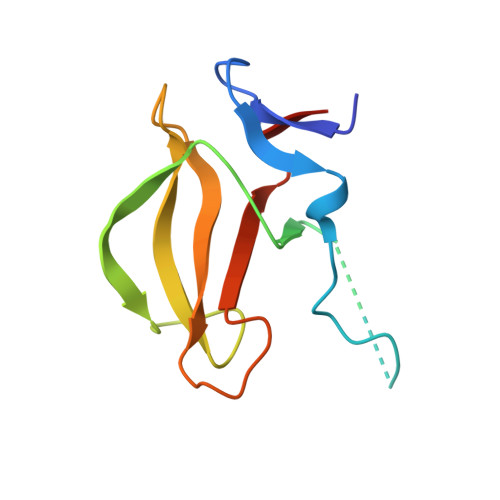
Structural basis of thalidomide enantiomer binding to cereblon
Mori, T., Ito, T., Liu, S., Ando, H., Sakamoto, S., Yamaguchi, Y., Tokunaga, E., Shibata, N., Handa, H., Hakoshima, T.(2018) Sci Rep 8: 1294-1294
- PubMed: 29358579 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-018-19202-7
- Primary Citation Related Structures:
5YIZ, 5YJ0, 5YJ1 - PubMed Abstract:
Thalidomide possesses two optical isomers which have been reported to exhibit different pharmacological and toxicological activities. However, the precise mechanism by which the two isomers exert their different activities remains poorly understood. Here, we present structural and biochemical studies of (S)- and (R)-enantiomers bound to the primary target of thalidomide, cereblon (CRBN). Our biochemical studies employed deuterium-substituted thalidomides to suppress optical isomer conversion, and established that the (S)-enantiomer exhibited ~10-fold stronger binding to CRBN and inhibition of self-ubiquitylation compared to the (R)-enantiomer. The crystal structures of the thalidomide-binding domain of CRBN bound to each enantiomer show that both enantiomers bind the tri-Trp pocket, although the bound form of the (S)-enantiomer exhibited a more relaxed glutarimide ring conformation. The (S)-enantiomer induced greater teratogenic effects on fins of zebrafish compared to the (R)-enantiomer. This study has established a mechanism by which thalidomide exerts its effects in a stereospecific manner at the atomic level.
- Structural Biology Laboratory, Nara Institute of Science and Technology, 8916-5 Takayama, Ikoma, Nara, 630-0192, Japan.
Organizational Affiliation: