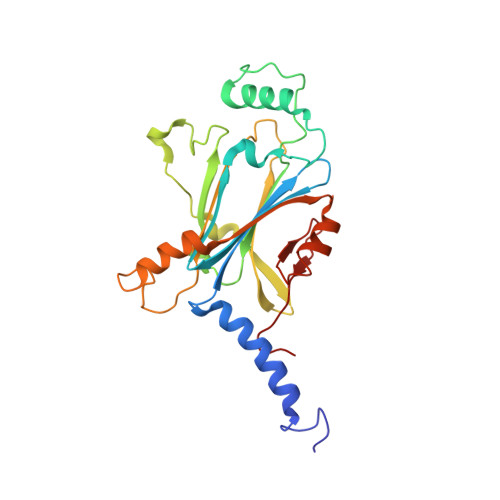
Structure of in cell protein crystals containing organometallic complexes.
Abe, S., Atsumi, K., Yamashita, K., Hirata, K., Mori, H., Ueno, T.(2018) Phys Chem Chem Phys 20: 2986-2989
- PubMed: 29138769 Search on PubMed
- DOI: https://doi.org/10.1039/c7cp06651a
- Primary Citation Related Structures:
5YHA, 5YHB - PubMed Abstract:
The molecular structures of in cell protein crystals containing organometallic Pd(allyl) complexes were determined by performing microfocus X-ray diffraction experiments. The coordination sites in a polyhedrin mutant with deletion of selected amino acid residues located at the interface of the polyhedrin trimer are dramatically altered compared to those of the wild-type composite.
- School of Life Science and Technology, Tokyo Institute of Technology, Nagatsuta-cho, Midori-ku, Yokohama 226-8501, Japan. saabe@bio.titech.ac.jp tueno@bio.titech.ac.jp.
Organizational Affiliation: