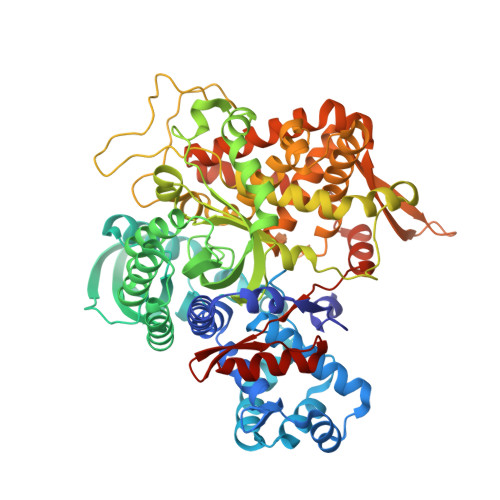
Crystal structures of Aflatoxin-oxidase from Armillariella tabescens reveal a dual activity enzyme.
Xu, T., Xie, C., Yao, D., Zhou, C.Z., Liu, J.(2017) Biochem Biophys Res Commun 494: 621-625
- PubMed: 29050944 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2017.10.077
- Primary Citation Related Structures:
5YFB, 5YFC, 5YFD - PubMed Abstract:
Aflatoxin-oxidase (AFO), a newly discovered oxidase isolated from Armillariella tabescens, was reported to perform aflatoxin B1 (AFB1) detoxification through breaking the bisfuran ring of AFB1. However, based on sequence alignment, we found that AFO shares high sequence identities with dipeptidyl peptidase III (DPP III) family members. To understand the functions of AFO, we determined its crystal structures in the absence and presence of zinc, copper ion, and employed HPLC to test if AFO could cleave the substrates of DPP III. Our structures reveal that AFO contains the classic DPP III activity center and the HPLC results further confirm that AFO possesses the dipeptidyl peptidase activity. Therefore, AFO should belong to DPP III family. Interestingly, unlike reported classic DPP III structure that has a large domain movement upon substrate binding, the AFO structures all adopt the closed conformation, independent of substrate binding. This conformation characteristic of AFO may be related to its enzyme activities. Taken together, our results demonstrate that AFO is a dual activity enzyme with both aflatoxin-oxidase and dipeptidyl peptidase activities and its unique conformation feature expands our understanding on the mode of reaction for this enzyme family.
- School of Life Sciences, University of Science and Technology of China, Hefei, 230026, China; State Key Laboratory of Respiratory Disease, Guangzhou Institutes of Biomedicine and Health, Chinese Academy of Sciences, Guangzhou, 510530, China; Guangdong Provincial Key Laboratory of Biocomputing, Guangzhou Institutes of Biomedicine and Health, Chinese Academy of Sciences, Guangzhou, 510530, China.
Organizational Affiliation: