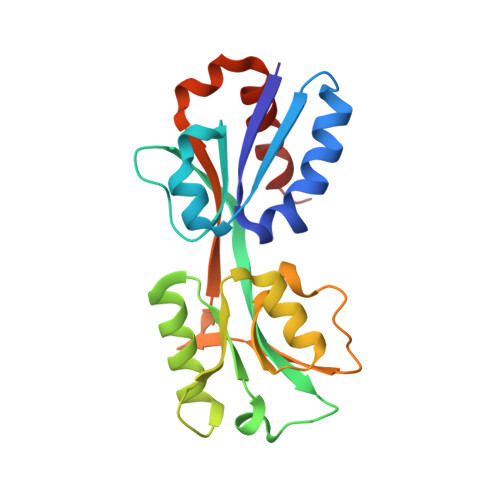
Structural and Biochemical Analysis of the Citrate-Responsive Mechanism of the Regulatory Domain of Catabolite Control Protein E from Staphylococcus aureus
Chen, J., Shang, F., Wang, L., Zou, L., Bu, T., Jin, L., Dong, Y., Ha, N.C., Quan, C., Nam, K.H., Xu, Y.(2018) Biochemistry 57: 6054-6060
- PubMed: 30252448 Search on PubMed
- DOI: https://doi.org/10.1021/acs.biochem.8b00671
- Primary Citation Related Structures:
5Y9Q, 5ZZO - PubMed Abstract:
Catabolite control protein E (CcpE) is a LysR-type transcriptional regulator that positively regulates the transcription of the first two enzymes of the TCA cycle, namely, citZ and citB, by sensing accumulated intracellular citrate. CcpE comprises an N-terminal DNA-binding domain and a C-terminal regulatory domain (RD) and senses citrate with conserved arginine residues in the RD. Although the crystal structure of the apo SaCcpE-RD has been reported, the citrate-responsive and DNA-binding mechanisms by which CcpE regulates TCA activity remain unclear. Here, we report the crystal structure of the apo and citrate-bound SaCcpE-RDs. The SaCcpE-RD exhibits conformational changes between the two subdomains via hinge motion of the central β4 and β10 strands. The citrate molecule is located in a positively charged cavity between the two subdomains and interacts with the highly conserved Ser98, Leu100, Arg145, and Arg256 residues. Compared with that of the apo SaCcpE-RD, the distance between the two subdomains of the citrate-bound SaCcpE-RD is more than ∼3 Å due to the binding of the citrate molecule, and this form exhibits a closed structure. The SaCcpE-RD exhibits various citrate-binding-independent conformational changes at the contacting interface. The SaCcpE-RD prefers the dimeric state in solution, whereas the SaCcpE-FL prefers the tetrameric state. Our results provide insight into the molecular function of SaCcpE.
- Department of Bioengineering, College of Life Science , Dalian Minzu University , Dalian 116600 , Liaoning , China.
Organizational Affiliation: