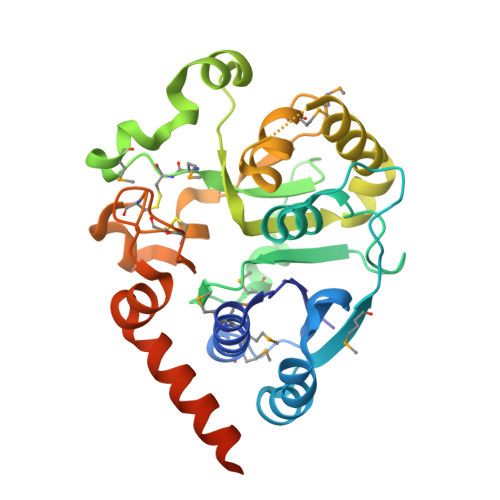
Visualisation of a flexible modular structure of the ER folding-sensor enzyme UGGT.
Satoh, T., Song, C., Zhu, T., Toshimori, T., Murata, K., Hayashi, Y., Kamikubo, H., Uchihashi, T., Kato, K.(2017) Sci Rep 7: 12142-12142
- PubMed: 28939828 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-017-12283-w
- Primary Citation Related Structures:
5H18, 5Y7F, 5Y7O - PubMed Abstract:
In the endoplasmic reticulum (ER), a protein quality control system facilitates the efficient folding of newly synthesised proteins. In this system, a series of N-linked glycan intermediates displayed on the protein surface serve as quality tags. The ER folding-sensor enzyme UDP-glucose:glycoprotein glucosyltransferase (UGGT) acts as a gatekeeper in the ER quality control system by specifically catalysing monoglucosylation onto incompletely folded glycoproteins, thereby enabling them to interact with lectin-chaperone complexes. Here we characterise the dynamic structure of this enzyme. Our crystallographic data demonstrate that the sensor region is composed of four thioredoxin-like domains followed by a β-rich domain, which are arranged into a C-shaped structure with a large central cavity, while the C-terminal catalytic domain undergoes a ligand-dependent conformational alteration. Furthermore, small-angle X-ray scattering, cryo-electron microscopy and high-speed atomic force microscopy have demonstrated that UGGT has a flexible modular structure in which the smaller catalytic domain is tethered to the larger folding-sensor region with variable spatial arrangements. These findings provide structural insights into the working mechanism whereby UGGT operates as a folding-sensor against a variety of glycoprotein substrates through its flexible modular structure possessing extended hydrophobic surfaces for the recognition of unfolded substrates.
- Graduate School of Pharmaceutical Sciences, Nagoya City University, 3-1 Tanabe-dori, Mizuho-ku, Nagoya, 467-8603, Japan. tadashisatoh@phar.nagoya-cu.ac.jp.
Organizational Affiliation: