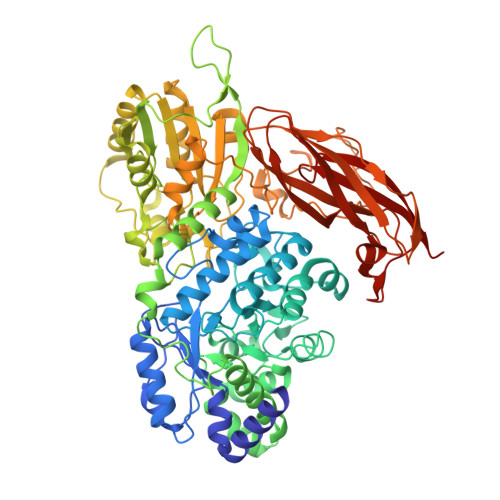
Function and structure relationships of a beta-1,2-glucooligosaccharide-degrading beta-glucosidase.
Ishiguro, R., Tanaka, N., Abe, K., Nakajima, M., Maeda, T., Miyanaga, A., Takahashi, Y., Sugimoto, N., Nakai, H., Taguchi, H.(2017) FEBS Lett 591: 3926-3936
- PubMed: 29131329 Search on PubMed
- DOI: https://doi.org/10.1002/1873-3468.12911
- Primary Citation Related Structures:
5XXL, 5XXM, 5XXN, 5XXO - PubMed Abstract:
BT_3567 protein, a putative β-glucosidase from Bacteroides thetaiotaomicron, exhibits higher activity toward Sop 3-5 (Sop n , n: degree of polymerization of β-1,2-glucooligosaccharides) than toward Sop 2 , unlike a known β-glucosidase from Listeria innocua which predominantly prefers Sop 2 . In the complex structure determined by soaking of a D286N mutant crystal with Sop 4 , a Sop 3 moiety was observed at subsites -1 to +2. The glucose moiety at subsite +2 forms a hydrogen bond with Asn81, which is replaced with Gly in the L. innocua β-glucosidase. The K m values of the N81G mutant for Sop 3-5 are much higher than those of the wild-type, suggesting that Asn81 contributes to the binding to substrates longer than Sop 3 .
- Department of Applied Biological Science, Faculty of Science and Technology, Tokyo University of Science, Noda, Chiba, Japan.
Organizational Affiliation: