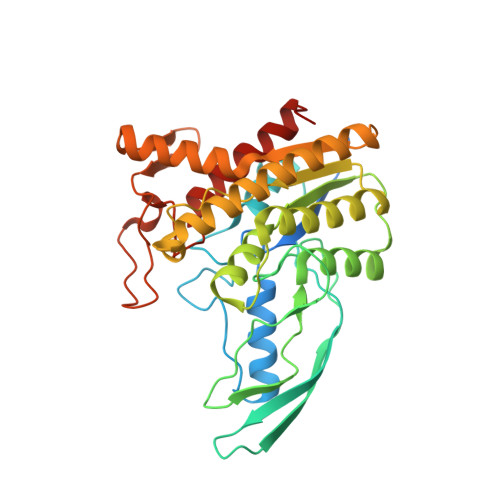
Crystallization and structure elucidation of GDSL esterase of Photobacterium sp. J15.
Mazlan, S.N.H.S., Ali, M.S.M., Rahman, R.N.Z.R.A., Sabri, S., Jonet, M.A., Leow, T.C.(2018) Int J Biol Macromol 119: 1188-1194
- PubMed: 30102982 Search on PubMed
- DOI: https://doi.org/10.1016/j.ijbiomac.2018.08.022
- Primary Citation Related Structures:
5XTU - PubMed Abstract:
GDSL esterase J15 (EstJ15) is a member of Family II of lipolytic enzyme. The enzyme was further classified in subgroup SGNH hydrolase due to the presence of highly conserve motif, Ser-Gly-Asn-His in four conserved blocks I, II, III, and V, respectively. X-ray quality crystal of EstJ15 was obtained from optimized formulation containing 0.10 M ammonium sulphate, 0.15 M sodium cacodylate trihydrate pH 6.5, and 20% PEG 8000. The crystal structure of EstJ15 was solved at 1.38 Å with one molecule per asymmetric unit. The structure exhibits α/β hydrolase fold and shared low amino acid sequence identity of 23% with the passenger domain of the autotransporter EstA of Pseudomonas aeruginosa. The active site is located at the centre of the structure, formed a narrow tunnel that hinder long substrates to be catalysed which was proven by the protein-ligand docking analysis. This study facilitates the understanding of high substrate specificity of EstJ15 and provide insights on its catalytic mechanism.
- Department of Cell and Molecular Biology, Faculty of Biotechnology and Biomolecular Sciences, Universiti Putra Malaysia, 43400 UPM Serdang, Selangor, Malaysia; Enzyme and Microbial Technology Research Center, Malaysia.
Organizational Affiliation: