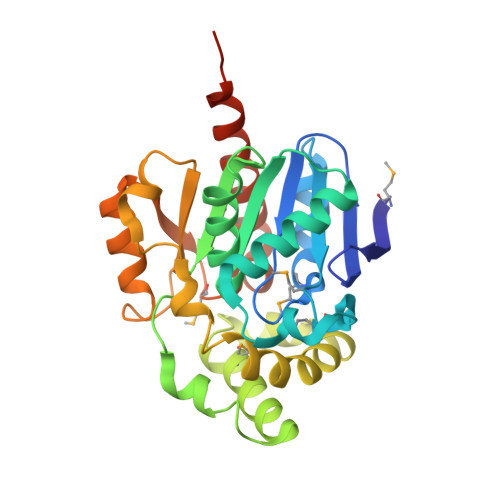
The structure of a complex of the lactonohydrolase zearalenone hydrolase with the hydrolysis product of zearalenone at 1.60 angstrom resolution
Qi, Q., Yang, W.J., Zhou, H.J., Ming, D.M., Sun, K.L., Xu, T.Y., Hu, X.J., Lv, H.(2017) Acta Crystallogr F Struct Biol Commun 73: 376-381
- PubMed: 28695844 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X17007713
- Primary Citation Related Structures:
5C8Z, 5XMW - PubMed Abstract:
Zearalenone hydrolase (ZHD) is an α/β-hydrolase that detoxifies and degrades the lactone zearalenone (ZEN), a naturally occurring oestrogenic mycotoxin that contaminates crops. Several apoenzyme and enzyme-substrate complex structures have been reported in the resolution range 2.4-2.6 Å. However, the properties and mechanism of this enzyme are not yet fully understood. Here, a 1.60 Å resolution structure of a ZHD-product complex is reported which was determined from a C-terminally His 6 -tagged ZHD crystal soaked with 2 mM ZEN for 30 min. It shows that after the lactone-bond cleavage, the phenol-ring region moves closer to residues Leu132, Tyr187 and Pro188, while the lactone-ring region barely moves. Comparisons of the ZHD-substrate and ZHD-product structures show that the hydrophilic interactions change, especially Trp183 N ℇ1 , which shifts from contacting O2 to O12', suggesting that Trp183 is responsible for the unidirectional translational movement of the phenol ring. This structure provides information on the final stage of the catalytic mechanism of zearalenone hydrolysis.
- State Key Laboratory of Genetic Engineering, School of Life Sciences, Fudan University, Shanghai 200438, People's Republic of China.
Organizational Affiliation: