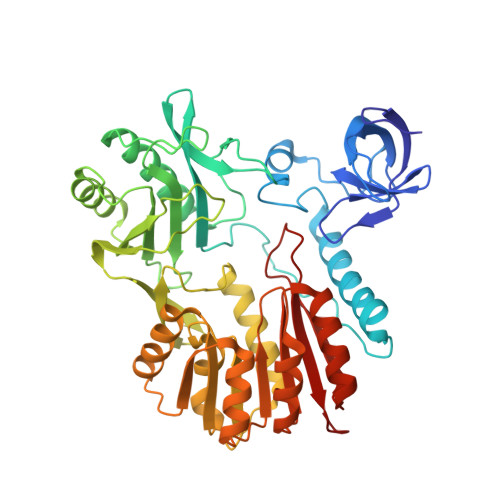
Structural insights into substrate selectivity of ribosomal RNA methyltransferase RlmCD
Jiang, Y., Li, F., Wu, J., Shi, Y., Gong, Q.(2017) PLoS One 12: e0185226-e0185226
- PubMed: 28949991 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0185226
- Primary Citation Related Structures:
5XJ1, 5XJ2 - PubMed Abstract:
RlmCD has recently been identified as the S-adenosyl methionine (SAM)-dependent methyltransferase responsible for the formation of m5U at U747 and U1939 of 23S ribosomal RNA in Streptococcus pneumoniae. In this research, we determine the high-resolution crystal structures of apo-form RlmCD and its complex with SAH. Using an in-vitro methyltransferase assay, we reveal the crucial residues for its catalytic functions. Furthermore, structural comparison between RlmCD and its structural homologue RumA, which only catalyzes the m5U1939 in Escherichia coli, implicates that a unique long linker in the central domain of RlmCD is the key factor in determining its substrate selectivity. Its significance in the enzyme activity of RlmCD is further confirmed by in-vitro methyltransferase assay.
- Hefei National Laboratory For Physical Sciences at Microscale and School of Life Sciences, University of Science and Technology of China, Hefei, Anhui, China.
Organizational Affiliation: