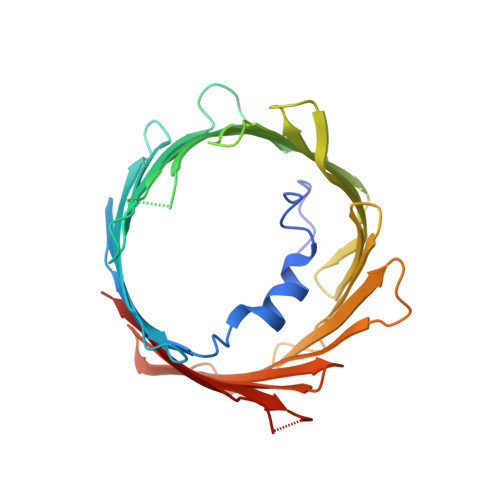
Crystal structural characterization reveals novel oligomeric interactions of human voltage-dependent anion channel 1
Hosaka, T., Okazaki, M., Kimura-Someya, T., Ishizuka-Katsura, Y., Ito, K., Yokoyama, S., Dodo, K., Sodeoka, M., Shirouzu, M.(2017) Protein Sci 26: 1749-1758
- PubMed: 28608415 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.3211
- Primary Citation Related Structures:
5XDN, 5XDO - PubMed Abstract:
Voltage-dependent anion channel 1 (VDAC1), which is located in the outer mitochondrial membrane, plays important roles in various cellular processes. For example, oligomerization of VDAC1 is involved in the release of cytochrome c to the cytoplasm, leading to apoptosis. However, it is unknown how VDAC1 oligomerization occurs in the membrane. In the present study, we determined high-resolution crystal structures of oligomeric human VDAC1 (hVDAC1) prepared by using an Escherichia coli cell-free protein synthesis system, which avoided the need for denaturation and refolding of the protein. Broad-range screening using a bicelle crystallization method produced crystals in space groups C222 and P22 1 2 1 , which diffracted to a resolution of 3.10 and 3.15 Å, respectively. Each crystal contained two hVDAC1 protomers in the asymmetric unit. Dimer within the asymmetrical unit of the crystal in space group C222 were oriented parallel, whereas those of the crystal in space group P22 1 2 1 were oriented anti-parallel. From a model of the crystal in space group C222, which we constructed by using crystal symmetry operators, a heptameric structure with eight patterns of interaction between protomers, including hydrophobic interactions with β-strands, hydrophilic interactions with loop regions, and protein-lipid interactions, was observed. It is possible that by having multiple patterns of interaction, VDAC1 can form homo- or hetero-oligomers not only with other VDAC1 protomers but also with other proteins such as VDAC2, VDAC3 and apoptosis-regulating proteins in the Bcl-2 family.
- Division of Structural and Synthetic Biology, RIKEN Center for Life Science Technologies, Yokohama, Kanagawa, 230-0045, Japan.
Organizational Affiliation: