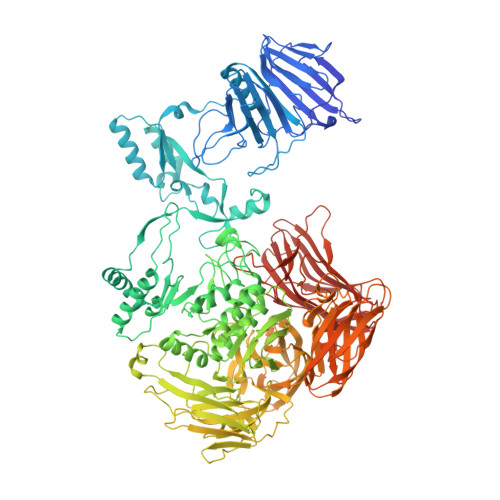
Carbohydrate-binding architecture of the multi-modular alpha-1,6-glucosyltransferase from Paenibacillus sp. 598K, which produces alpha-1,6-glucosyl-alpha-glucosaccharides from starch
Fujimoto, Z., Suzuki, N., Kishine, N., Ichinose, H., Momma, M., Kimura, A., Funane, K.(2017) Biochem J 474: 2763-2778
- PubMed: 28698247 Search on PubMed
- DOI: https://doi.org/10.1042/BCJ20170152
- Primary Citation Related Structures:
5X7O, 5X7P, 5X7Q, 5X7R, 5X7S - PubMed Abstract:
Paenibacillus sp. 598K α-1,6-glucosyltransferase (Ps6TG31A), a member of glycoside hydrolase family 31, catalyzes exo-α-glucohydrolysis and transglucosylation and produces α-1,6-glucosyl-α-glucosaccharides from α-glucan via its disproportionation activity. The crystal structure of Ps6TG31A was determined by an anomalous dispersion method using a terbium derivative. The monomeric Ps6TG31A consisted of one catalytic (β/α) 8 -barrel domain and six small domains, one on the N-terminal and five on the C-terminal side. The structures of the enzyme complexed with maltohexaose, isomaltohexaose, and acarbose demonstrated that the ligands were observed in the catalytic cleft and the sugar-binding sites of four β-domains. The catalytic site was structured by a glucose-binding pocket and an aglycon-binding cleft built by two sidewalls. The bound acarbose was located with its non-reducing end pseudosugar docked in the pocket, and the other moieties along one sidewall serving three subsites for the α-1,4-glucan. The bound isomaltooligosaccharide was found on the opposite sidewall, which provided the space for the acceptor molecule to be positioned for attack of the catalytic intermediate covalent complex during transglucosylation. The N-terminal domain recognized the α-1,4-glucan in a surface-binding mode. Two of the five C-terminal domains belong to the carbohydrate-binding modules family 35 and one to family 61. The sugar complex structures indicated that the first family 35 module preferred α-1,6-glucan, whereas the second family 35 module and family 61 module preferred α-1,4-glucan. Ps6TG31A appears to have enhanced transglucosylation activity facilitated by its carbohydrate-binding modules and substrate-binding cleft that positions the substrate and acceptor sugar for the transglucosylation.
- Advanced Analysis Center, National Agriculture and Food Research Organization (NARO), 2-1-2 Kannondai, Tsukuba, Ibaraki 305-8602, Japan zui@affrc.go.jp funane@affrc.go.jp.
Organizational Affiliation: