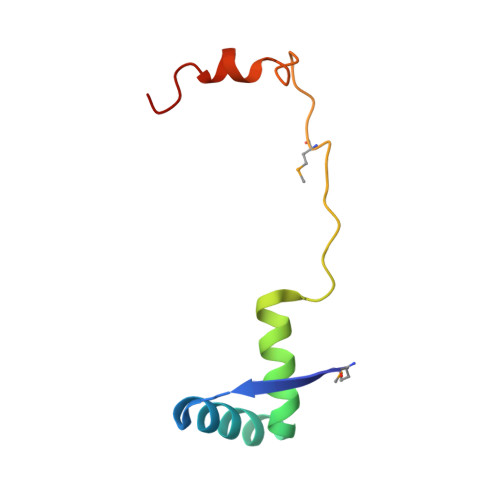
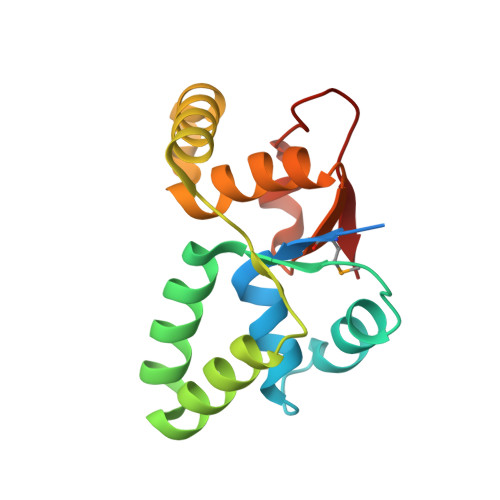
Functional details of the Mycobacterium tuberculosis VapBC26 toxin-antitoxin system based on a structural study: insights into unique binding and antibiotic peptides.
Kang, S.M., Kim, D.H., Lee, K.Y., Park, S.J., Yoon, H.J., Lee, S.J., Im, H., Lee, B.J.(2017) Nucleic Acids Res 45: 8564-8580
- PubMed: 28575388 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkx489
- Primary Citation Related Structures:
5X3T - PubMed Abstract:
Toxin-antitoxin (TA) systems are essential for bacterial persistence under stressful conditions. In particular, Mycobacterium tuberculosis express VapBC TA genes that encode the stable VapC toxin and the labile VapB antitoxin. Under normal conditions, these proteins interact to form a non-toxic TA complex, but the toxin is activated by release from the antitoxin in response to unfavorable conditions. Here, we present the crystal structure of the M. tuberculosis VapBC26 complex and show that the VapC26 toxin contains a pilus retraction protein (PilT) N-terminal (PIN) domain that is essential for ribonuclease activity and that, the VapB26 antitoxin folds into a ribbon-helix-helix DNA-binding motif at the N-terminus. The active site of VapC26 is sterically blocked by the flexible C-terminal region of VapB26. The C-terminal region of free VapB26 adopts an unfolded conformation but forms a helix upon binding to VapC26. The results of RNase activity assays show that Mg2+ and Mn2+ are essential for the ribonuclease activity of VapC26. As shown in the nuclear magnetic resonance spectra, several residues of VapB26 participate in the specific binding to the promoter region of the VapBC26 operon. In addition, toxin-mimicking peptides were designed that inhibit TA complex formation and thereby increase toxin activity, providing a novel approach to the development of new antibiotics.
- The Research Institute of Pharmaceutical Sciences, College of Pharmacy, Seoul National University, Gwanak-gu, Seoul 151-742, Republic of Korea.
Organizational Affiliation: