Cleavage of peptidic inhibitors by target protease is caused by peptide conformational transition.
Jiang, L., Oldenburg, E., Kromann-Hansen, T., Xu, P., Jensen, J.K., Andreasen, P.A., Huang, M.(2018) Biochim Biophys Acta 1862: 2017-2023
- PubMed: 29959058 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbagen.2018.06.016
- Primary Citation Related Structures:
5WXF, 5WXO, 5WXP - PubMed Abstract:
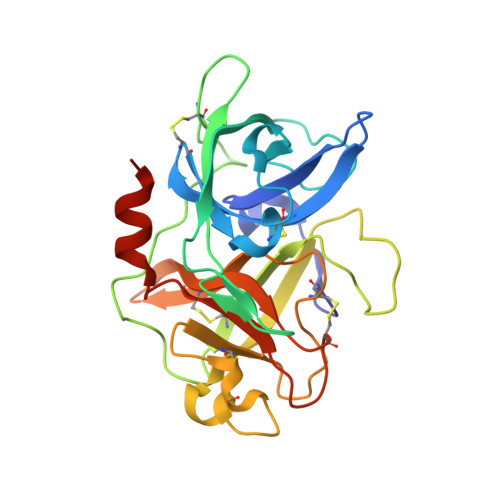
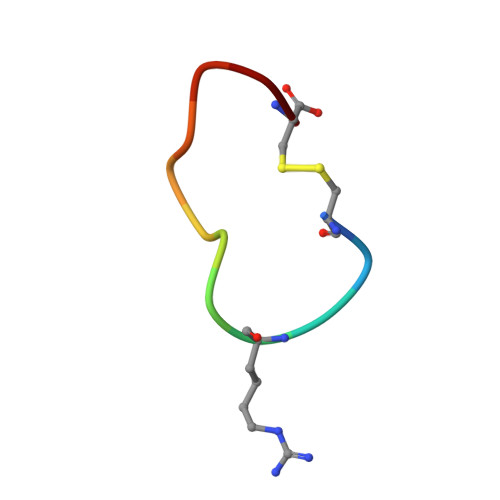
Some peptide sequences can behave as either substrates or inhibitors of serine proteases. Working with a cyclic peptidic inhibitor of the serine protease urokinase-type plasminogen activator (uPA), we have now demonstrated a new mechanism for an inhibitor-to-substrate switch. The peptide, CSWRGLENHAAC (upain-2), is a competitive inhibitor of human uPA, but is also slowly converted to a substrate in which the bond between Arg 4 and Gly 5 (the P1-P1' bond) is cleaved. Substituting the P2 residue Trp 3 to an Ala or substituting the P1 Arg 4 residue with 4-guanidino-phenylalanine strongly increased the substrate cleavage rate. We studied the structural basis for the inhibitor-to-substrate switch by determining the crystal structures of the various peptide variants in complex with the catalytic domain of uPA. While the slowly cleaved peptides bound clearly in inhibitory mode, with the oxyanion hole blocked by the side chain of the P3' residue Glu 7 , peptides behaving essentially as substrates with a much accelerated rate of cleavage was observed to be bound to the enzyme in substrate mode. Our analysis reveals that the inhibitor-to-substrate switch was associated with a 7 Å translocation of the P2 residue, and we conclude that the inhibitor-to-substrate switch of upain-2 is a result of a major conformational change in the enzyme-bound state of the peptide. This conclusion is in contrast to findings with so-called standard mechanism inhibitors in which the inhibitor-to-substrate switch is linked to minor conformational changes in the backbone of the inhibitory peptide stretch.
- College of Chemistry, Fuzhou University, 350116, China.
Organizational Affiliation: