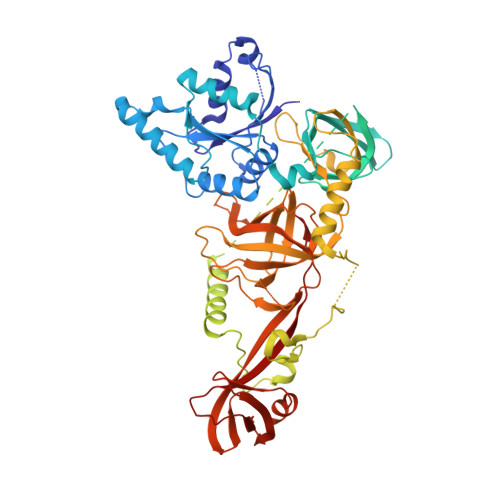
Molecular architecture of the 90S small subunit pre-ribosome
Sun, Q., Zhu, X., Qi, J., An, W., Lan, P., Tan, D., Chen, R., Wang, B., Zheng, S., Zhang, C., Chen, X., Zhang, W., Chen, J., Dong, M.Q., Ye, K.(2017) Elife 6
- PubMed: 28244370 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.22086
- Primary Citation Related Structures:
5WWN, 5WWO, 5WXL, 5WXM, 5WY3, 5WYJ, 5WYK, 5WYL - PubMed Abstract:
Eukaryotic small ribosomal subunits are first assembled into 90S pre-ribosomes. The complete 90S is a gigantic complex with a molecular mass of approximately five megadaltons. Here, we report the nearly complete architecture of Saccharomyces cerevisiae 90S determined from three cryo-electron microscopy single particle reconstructions at 4.5 to 8.7 angstrom resolution. The majority of the density maps were modeled and assigned to specific RNA and protein components. The nascent ribosome is assembled into isolated native-like substructures that are stabilized by abundant assembly factors. The 5' external transcribed spacer and U3 snoRNA nucleate a large subcomplex that scaffolds the nascent ribosome. U3 binds four sites of pre-rRNA, including a novel site on helix 27 but not the 3' side of the central pseudoknot, and crucially organizes the 90S structure. The 90S model provides significant insight into the principle of small subunit assembly and the function of assembly factors.
- PTN Joint Graduate Program, School of Life Sciences, Tsinghua University, Beijing, China.
Organizational Affiliation: