Green fluorescent protein linked peptide inhibitor PMI in complex with mdm2
Chee, S.M.Q., Wongsantichon, J., Sana, B., Robinson, R.C., Ghadessy, F.J.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Green fluorescent protein linked MTide-02 | 358 | Aequorea victoria, Camelus dromedarius This entity is chimeric | Mutation(s): 0 Gene Names: GFP | 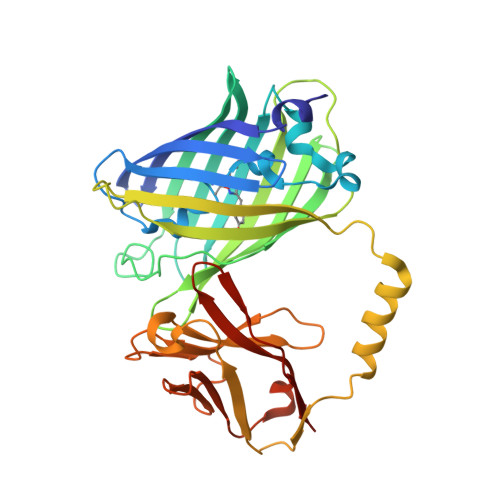 | |
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| E3 ubiquitin-protein ligase Mdm2 | 122 | Homo sapiens | Mutation(s): 0 Gene Names: MDM2 EC: 2.3.2.27 | 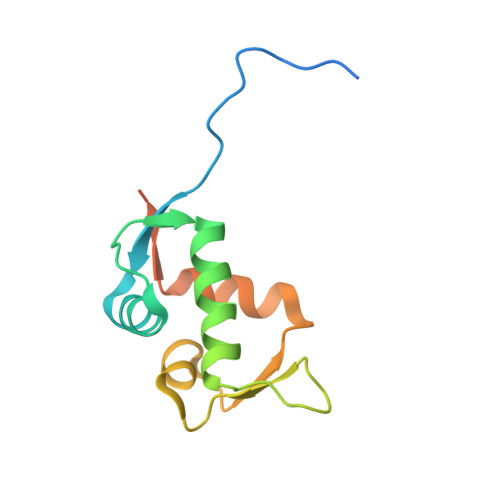 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q00987 GTEx: ENSG00000135679 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q00987 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| PG4 Download:Ideal Coordinates CCD File | E [auth B] | TETRAETHYLENE GLYCOL C8 H18 O5 UWHCKJMYHZGTIT-UHFFFAOYSA-N |  | ||
| PO4 Download:Ideal Coordinates CCD File | D [auth A], F [auth B] | PHOSPHATE ION O4 P NBIIXXVUZAFLBC-UHFFFAOYSA-K |  | ||
| GOL Download:Ideal Coordinates CCD File | C [auth A] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| Modified Residues 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| CRO Query on CRO | A | L-PEPTIDE LINKING | C15 H17 N3 O5 |  | THR, TYR, GLY |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 51.766 | α = 90 |
| b = 98.422 | β = 90 |
| c = 218.988 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| HKL-2000 | data reduction |
| HKL-2000 | data scaling |
| PHASER | phasing |