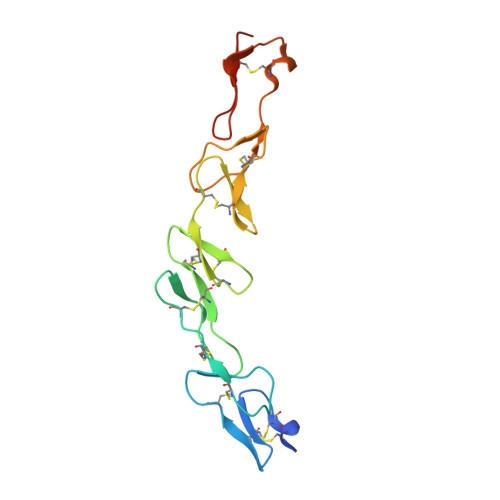
Crystal structure of murine 4-1BB and its interaction with 4-1BBL support a role for galectin-9 in 4-1BB signaling.
Bitra, A., Doukov, T., Wang, J., Picarda, G., Benedict, C.A., Croft, M., Zajonc, D.M.(2018) J Biological Chem 293: 1317-1329
- PubMed: 29242193 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M117.814905
- Primary Citation Related Structures:
5WI8, 5WIW, 5WJF - PubMed Abstract:
4-1BB (CD137) is a TNF receptor superfamily (TNFRSF) member that is thought to undergo receptor trimerization upon binding to its trimeric TNF superfamily ligand (4-1BBL) to stimulate immune responses. 4-1BB also can bind to the tandem repeat-type lectin galectin-9 (Gal-9), and signaling through mouse (m)4-1BB is reduced in galectin-9 (Gal-9)-deficient mice, suggesting a pivotal role of Gal-9 in m4-1BB activation. Here, using sulfur-SAD phasing, we determined the crystal structure of m4-1BB to 2.2-Å resolution. We found that similar to other TNFRSFs, m4-1BB has four cysteine-rich domains (CRDs). However, the organization of CRD1 and the orientation of CRD3 and CRD4 with respect to CRD2 in the m4-1BB structure distinctly differed from those of other TNFRSFs. Moreover, we mapped two Asn residues within CRD4 that are N -linked glycosylated and mediate m4-1BB binding to Gal-9. Kinetics studies of m4-1BB disclosed a very tight nanomolar binding affinity to m4-1BBL with an unexpectedly strong avidity effect. Both N- and C-terminal domains of Gal-9 bound m4-1BB, but with lower affinity compared with m4-1BBL. Although the TNF homology domain (THD) of human (h)4-1BBL forms non-covalent trimers, we found that m4-1BBL formed a covalent dimer via 2 cysteines absent in h4-1BBL. As multimerization and clustering is a prerequisite for TNFR intracellular signaling, and as m4-1BBL can only recruit two m4-1BB monomers, we hypothesize that m4-1BBL and Gal-9 act together to aid aggregation of m4-1BB monomers to efficiently initiate m4-1BB signaling.
- From the Division of Immune Regulation, La Jolla Institute for Allergy and Immunology (LJI), La Jolla, California 92037.
Organizational Affiliation: