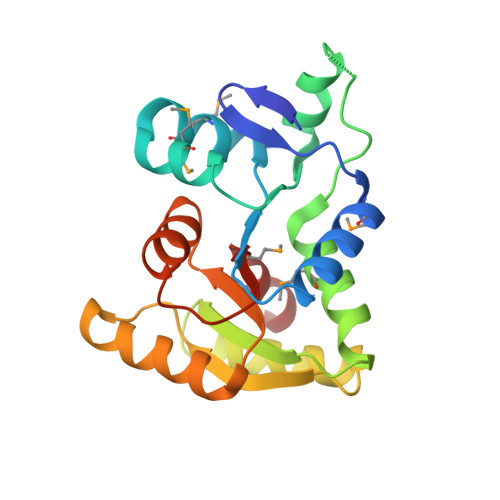
Structure and mechanism of TagA, a novel membrane-associated glycosyltransferase that produces wall teichoic acids in pathogenic bacteria.
Kattke, M.D., Gosschalk, J.E., Martinez, O.E., Kumar, G., Gale, R.T., Cascio, D., Sawaya, M.R., Philips, M., Brown, E.D., Clubb, R.T.(2019) PLoS Pathog 15: e1007723-e1007723
- PubMed: 31002736 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1007723
- Primary Citation Related Structures:
5WB4, 5WFG - PubMed Abstract:
Staphylococcus aureus and other bacterial pathogens affix wall teichoic acids (WTAs) to their surface. These highly abundant anionic glycopolymers have critical functions in bacterial physiology and their susceptibility to β-lactam antibiotics. The membrane-associated TagA glycosyltransferase (GT) catalyzes the first-committed step in WTA biosynthesis and is a founding member of the WecB/TagA/CpsF GT family, more than 6,000 enzymes that synthesize a range of extracellular polysaccharides through a poorly understood mechanism. Crystal structures of TagA from T. italicus in its apo- and UDP-bound states reveal a novel GT fold, and coupled with biochemical and cellular data define the mechanism of catalysis. We propose that enzyme activity is regulated by interactions with the bilayer, which trigger a structural change that facilitates proper active site formation and recognition of the enzyme's lipid-linked substrate. These findings inform upon the molecular basis of WecB/TagA/CpsF activity and could guide the development of new anti-microbial drugs.
- Department of Chemistry and Biochemistry, University of California, Los Angeles, Los Angeles, United States of America.
Organizational Affiliation: