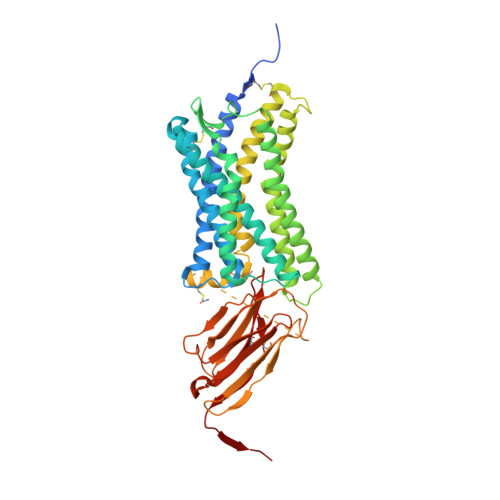
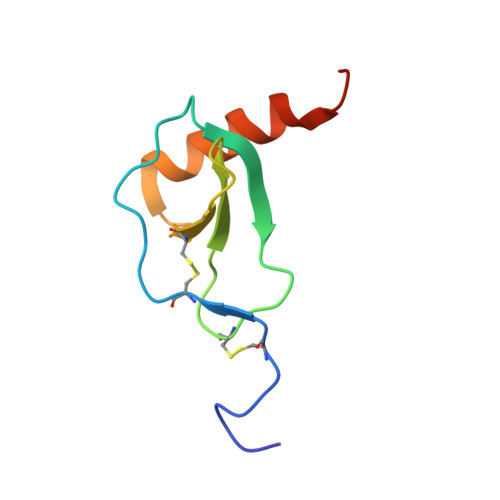
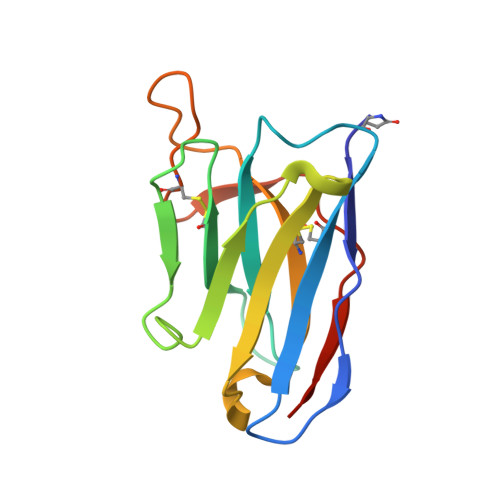
Viral GPCR US28 can signal in response to chemokine agonists of nearly unlimited structural degeneracy.
Miles, T.F., Spiess, K., Jude, K.M., Tsutsumi, N., Burg, J.S., Ingram, J.R., Waghray, D., Hjorto, G.M., Larsen, O., Ploegh, H.L., Rosenkilde, M.M., Garcia, K.C.(2018) Elife 7
- PubMed: 29882741 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.35850
- Primary Citation Related Structures:
5WB1, 5WB2 - PubMed Abstract:
Human cytomegalovirus has hijacked and evolved a human G-protein-coupled receptor into US28, which functions as a promiscuous chemokine 'sink' to facilitate evasion of host immune responses. To probe the molecular basis of US28's unique ligand cross-reactivity, we deep-sequenced CX3CL1 chemokine libraries selected on 'molecular casts' of the US28 active-state and find that US28 can engage thousands of distinct chemokine sequences, many of which elicit diverse signaling outcomes. The structure of a G-protein-biased CX3CL1-variant in complex with US28 revealed an entirely unique chemokine amino terminal peptide conformation and remodeled constellation of receptor-ligand interactions. Receptor signaling, however, is remarkably robust to mutational disruption of these interactions. Thus, US28 accommodates and functionally discriminates amongst highly degenerate chemokine sequences by sensing the steric bulk of the ligands, which distort both receptor extracellular loops and the walls of the ligand binding pocket to varying degrees, rather than requiring sequence-specific bonding chemistries for recognition and signaling.
- Department of Molecular and Cellular Physiology, Stanford University School of Medicine, Stanford, United States.
Organizational Affiliation: