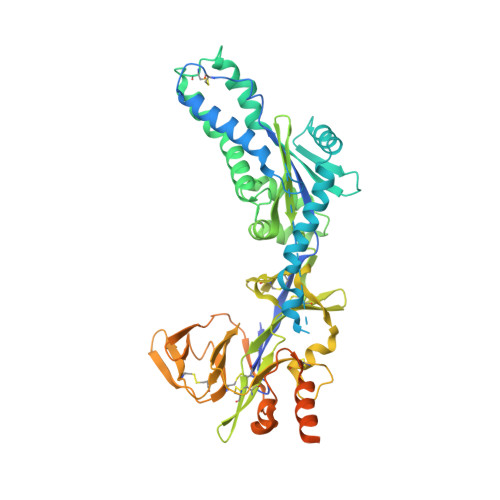
Structure and immunogenicity of pre-fusion-stabilized human metapneumovirus F glycoprotein.
Battles, M.B., Mas, V., Olmedillas, E., Cano, O., Vazquez, M., Rodriguez, L., Melero, J.A., McLellan, J.S.(2017) Nat Commun 8: 1528-1528
- PubMed: 29142300 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-017-01708-9
- Primary Citation Related Structures:
5WB0 - PubMed Abstract:
Human metapneumovirus (hMPV) is a frequent cause of bronchiolitis in young children. Its F glycoprotein mediates virus-cell membrane fusion and is the primary target of neutralizing antibodies. The inability to produce recombinant hMPV F glycoprotein in the metastable pre-fusion conformation has hindered structural and immunological studies. Here, we engineer a pre-fusion-stabilized hMPV F ectodomain and determine its crystal structure to 2.6 Å resolution. This structure reveals molecular determinants of strain-dependent acid-induced fusion, as well as insights into refolding from pre- to post-fusion conformations. A dense glycan shield at the apex of pre-fusion hMPV F suggests that antibodies against this site may not be elicited by host immune responses, which is confirmed by depletion studies of human immunoglobulins and by mouse immunizations. This is a major difference with pre-fusion F from human respiratory syncytial virus (hRSV), and collectively our results should facilitate development of effective hMPV vaccine candidates.
- Department of Biochemistry and Cell Biology, Geisel School of Medicine at Dartmouth, Hanover, NH, 03755, USA.
Organizational Affiliation: